Difference between revisions of "Team:DTU-Denmark/Journal"
| Line 5: | Line 5: | ||
| − | <link href="/Team:DTU-Denmark/ | + | <link href="/Team:DTU-Denmark/loadphpcss?action=raw&ctype=text/css" rel="stylesheet"> |
| − | <link href="/Team:DTU-Denmark/font- | + | <link href="/Team:DTU-Denmark/bootstrapcss?action=raw&ctype=text/css" rel="stylesheet"> |
| − | <link href="/Team:DTU-Denmark/ | + | <link href="/Team:DTU-Denmark/font-awesomecss?action=raw&ctype=text/css" rel="stylesheet"> |
| − | <link href="/Team:DTU-Denmark/ | + | <link href="/Team:DTU-Denmark/wikicss?action=raw&ctype=text/css" rel="stylesheet"> |
| − | <link href="/Team:DTU-Denmark/ | + | <link href="/Team:DTU-Denmark/timelinecss?action=raw&ctype=text/css" rel="stylesheet"> |
| + | <link href="/Team:DTU-Denmark/menufixcss?action=raw&ctype=text/css" rel="stylesheet"> | ||
| − | <script type="text/javascript" src="/Team:DTU-Denmark/ | + | <script type="text/javascript" src="/Team:DTU-Denmark/jqueryjs?action=raw&ctype=text/javascript"></script> |
| − | <script type="text/javascript" src="/Team:DTU-Denmark/ | + | <script type="text/javascript" src="/Team:DTU-Denmark/bootstrapjs?action=raw&ctype=text/javascript"></script> |
| − | <script type="text/javascript" src="/Team:DTU-Denmark/ | + | <script type="text/javascript" src="/Team:DTU-Denmark/jqueryeasingjs?action=raw&ctype=text/javascript"></script> |
| − | <script type="text/javascript" src="/Team:DTU-Denmark/ | + | <script type="text/javascript" src="/Team:DTU-Denmark/wikijs?action=raw&ctype=text/javascript"></script> |
<!-- MathJax (LaTeX for the web) --> | <!-- MathJax (LaTeX for the web) --> | ||
<script src="https://2015.igem.org/common/MathJax-2.5-latest/MathJax.js?config=TeX-AMS-MML_SVG"></script> | <script src="https://2015.igem.org/common/MathJax-2.5-latest/MathJax.js?config=TeX-AMS-MML_SVG"></script> | ||
| Line 37: | Line 38: | ||
<nav class="navbar navbar-default navbar-fixed-top" role="navigation"> | <nav class="navbar navbar-default navbar-fixed-top" role="navigation"> | ||
<!-- Brand and toggle get grouped for better mobile display --> | <!-- Brand and toggle get grouped for better mobile display --> | ||
| − | <div class="container"> | + | <div class="container-fluid"> |
<div class="navbar-header"> | <div class="navbar-header"> | ||
<button type="button" class="navbar-toggle" data-toggle="collapse" data-target="#nav-collapse"> | <button type="button" class="navbar-toggle" data-toggle="collapse" data-target="#nav-collapse"> | ||
<span class="sr-only">Toggle navigation</span> | <span class="sr-only">Toggle navigation</span> | ||
<i class="fa fa-bars"></i> | <i class="fa fa-bars"></i> | ||
| − | |||
| − | |||
| − | |||
</button> | </button> | ||
<a class="navbar-brand navimg" href="#"><img src="/wiki/images/b/b7/DTU-Denmark_dtulogo.png"></a> | <a class="navbar-brand navimg" href="#"><img src="/wiki/images/b/b7/DTU-Denmark_dtulogo.png"></a> | ||
</div> | </div> | ||
| − | |||
<!-- Collect the nav links, forms, and other content for toggling --> | <!-- Collect the nav links, forms, and other content for toggling --> | ||
<div class="collapse navbar-collapse" id="nav-collapse"> | <div class="collapse navbar-collapse" id="nav-collapse"> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<ul class="nav nav-justified"> | <ul class="nav nav-justified"> | ||
| Line 89: | Line 67: | ||
<li > | <li > | ||
| − | <a class="active" href="/Team:DTU-Denmark/ | + | <a class="active" href="/Team:DTU-Denmark/Part_Collection">Parts</a> |
</li> | </li> | ||
| Line 96: | Line 74: | ||
</li> | </li> | ||
| − | + | <li > | |
| − | + | <a class="active" href="/Team:DTU-Denmark/Software">Software</a> | |
| − | <li> | + | </li> |
| + | |||
| + | <li > | ||
| + | <a class="active" href="/Team:DTU-Denmark/Achivements">Achievements</a> | ||
| + | </li> | ||
| + | |||
| + | <li class="hidden-xs"> | ||
| + | <a href="http://facebook.com/dtubiobuilders" data-toggle="tooltip" data-placement="bottom" title="Follow us on Facebook"> | ||
| + | <i class="fa fa-facebook"></i><span class="visible-xs"> Follow us on Facebook</span> | ||
| + | </a> | ||
| + | </li> | ||
| + | <li class="visible-xs"><a href="http://facebook.com/dtubiobuilders"><i class="fa fa-facebook-square"></i> Follow us on Facebook</a></li> | ||
| + | <li class="hidden-xs"> | ||
| + | <a href="https://twitter.com/IGEM_DTU" data-toggle="tooltip" data-placement="bottom" title="Follow us on Twitter"> | ||
| + | <i class="fa fa-twitter"></i> | ||
| + | </a> | ||
| + | </li> | ||
| + | <li class="visible-xs"><a href="https://twitter.com/IGEM_DTU"><i class="fa fa-twitter-square"></i> Follow us on Twitter</a></li> | ||
| + | <li class="hidden-lg"> | ||
<a href="https://igem.org" target="_blank" class="navimg"> | <a href="https://igem.org" target="_blank" class="navimg"> | ||
<img src="/wiki/images/a/af/DTU-Denmark_igemlogo.png"> | <img src="/wiki/images/a/af/DTU-Denmark_igemlogo.png"> | ||
</a> | </a> | ||
</li> | </li> | ||
| − | |||
| − | |||
</ul> | </ul> | ||
</div><!-- /.navbar-collapse --> | </div><!-- /.navbar-collapse --> | ||
| Line 110: | Line 104: | ||
</nav> | </nav> | ||
<nav class="navbar navbar-default extrabar hidden-xs hidden" data-spy="affix"> | <nav class="navbar navbar-default extrabar hidden-xs hidden" data-spy="affix"> | ||
| − | <div class="container" id="scrollspy"> | + | <div class="container-fluid" id="scrollspy"> |
<ul class="nav nav-justified"> | <ul class="nav nav-justified"> | ||
<li><a class="page-scroll" href="#Summary">Summary</a></li><li><a class="page-scroll" href="#Methods">Methods</a></li><li><a class="page-scroll" href="#Timeline">Timeline</a></li> | <li><a class="page-scroll" href="#Summary">Summary</a></li><li><a class="page-scroll" href="#Methods">Methods</a></li><li><a class="page-scroll" href="#Timeline">Timeline</a></li> | ||
| Line 122: | Line 116: | ||
</a> | </a> | ||
</div> | </div> | ||
| − | <div class="container"> | + | <div class="container-fluid"> |
<div class="row hero-text"> | <div class="row hero-text"> | ||
<div class="col-md-12"> | <div class="col-md-12"> | ||
| Line 180: | Line 174: | ||
Summary | Summary | ||
</h1> | </h1> | ||
| − | <p>For | + | <p>For OGRE (Oligo Recombineering) to be effective there is two genetic improvements that can be done. The first and most important is that the organism need to express a recombinase protein such as the beta protein derived from the lambda RED E. coli phage. The second improvement is to destroy/inhibit the mismatch repair system [1]. In B. subtilis the genes mutS and mutL are taking care of the mismatch repair [2]. For proof of concept purposes deleting the mismatch repair system will be adequate. According to (REF) the protein encoded by mutL will not be functional without a functional mutS, this is why deletion of only mutS will be sufficient to take out the mismatch repair system. With respect to recombinase we will test two different recombinases: lambda-beta (codon optimized for <em>B. subtilis 168</em>) and gp35 (a recombinase from the B. subtilis phage SPP1) [3]. For duing proof of concept of OGRE in <em>B. subtilis</em> four different strains was producere via genetic recombineering. All derived from <em>Bacillus subtilis 168</em>:</p> |
<ul> | <ul> | ||
<li>∆<em>amyE::beta-neo<sup>R</sup></em></li> | <li>∆<em>amyE::beta-neo<sup>R</sup></em></li> | ||
| − | <li>∆<em>amyE:: | + | <li>∆<em>amyE::GP35-neo<sup>R</sup></em></li> |
<li>∆<em>mutS::beta-neo<sup>R</sup></em></li> | <li>∆<em>mutS::beta-neo<sup>R</sup></em></li> | ||
| − | <li>∆<em>mutS:: | + | <li>∆<em>mutS::GP35-neo<sup>R</sup></em></li> |
</ul> | </ul> | ||
| + | |||
| + | <p>The four different strains where compared to each other to get compare there transformation efficienc in MAGE.</p> | ||
</div> | </div> | ||
| Line 200: | Line 196: | ||
Methods | Methods | ||
</h1> | </h1> | ||
| − | <p> | + | <p> </p> |
| − | <p><strong> | + | <p><strong>making beta and GP35</strong></p> |
| − | <p> | + | <p>The beta CDS was codon optimized with JCat() and ordered from COMPANY. GP35 sequence was found from (GP35 article). The sequence was ordered from the same company as beta.</p> |
| − | <p><strong> | + | <p><strong>incertion of recombinases in the amyE locus of B. subtilis</strong></p> |
| + | |||
| + | <p>Beta and GP35 was inserted into the plasmid known as pDG268neo by using PCR with G5 HF 2x master mix from NEB (primer sequence hyperlink?). The plasmid was transformed into E coli DH5alpha by heat chock and incubated over night. The plasmid was purified from E coli by using QiAPrep mini prep kit. Digesting with restriction enzymes validated the insertion. This plasmid integrates the CDS into the amyE locus in Bacillus subtilis. pDG268neo with ether beta or GP35 was linearized and transformed into B subtilis W168 using the CFrench protocol (make CFrench to hyperlink)</p> | ||
| + | |||
| + | <p>chromosomal DNA from B. subtilis 168 was isolated <strong>(protocol)</strong></p> | ||
| + | |||
| + | <p><strong>knock out mutS </strong></p> | ||
| + | |||
| + | <p>The regions upstream and downstream of the mutS locus were amplified by PCR from the chromosomal DNA with primers that had overlap with GP35 and beta.</p> | ||
| + | |||
| + | <p>GP35 and beta were PCR amplified with primers that had overhangs that were homologies to the mutS locus in bacillus subtilis. GP35 and beta was assembled into the pSB1C3 plasmid by using gibson assembly and transformed into E coli and incubated over night. The plasmid was isolated by using mini prep. The plasmids were verified using restriction enzymes.</p> | ||
| + | |||
| + | <p>beta and GP35 in pSB1C3 was linearized using XhoI, This also removed the Chloramphenicol gene from the plasmid. Beta and GP35 was transformed into B. subtilis.</p> | ||
| + | |||
| + | <p> | ||
| + | <p><strong>Genomic DNA extraction (B. brevis)</strong></p> | ||
| + | </p> | ||
| + | <p> | ||
<p><strong>Ligation of DNA</strong></p> | <p><strong>Ligation of DNA</strong></p> | ||
| + | </p> | ||
<p><strong>PCR</strong> -</p> | <p><strong>PCR</strong> -</p> | ||
| Line 224: | Line 238: | ||
<p><strong>Restriction enzyme digest (NEB)</strong></p> | <p><strong>Restriction enzyme digest (NEB)</strong></p> | ||
| + | |||
| + | <p><strong>Tyrocidine construct</strong></p> | ||
<p><strong>Transformation and Preparation of E.coli</strong><br /> | <p><strong>Transformation and Preparation of E.coli</strong><br /> | ||
| Line 229: | Line 245: | ||
-Transformation of Chemically Competent E.coli<br /> | -Transformation of Chemically Competent E.coli<br /> | ||
<strong>Transformation of Bacillus</strong></p> | <strong>Transformation of Bacillus</strong></p> | ||
| + | |||
| + | <p><strong>Optiminsing OGRE in Bacillus</strong></p> | ||
| + | |||
| + | <p style="margin-left:30.0pt;">Different amount of oligo was tested to optimize OGRE method for Bacillus subtilis.</p> | ||
| + | |||
| + | <p style="margin-left:30.0pt;">The mismatch insertion frequency and oligo length was tested. The standard OGRE protocol was follow with the mismatches verified from 1-6 mismatches and oligoes from 50-100nt.</p> | ||
| + | |||
| + | <p style="margin-left:30.0pt;">OGRE with 4 cycles were attempted every cycle to approximately 6 hours. This was done to test if the transformation efficiency would rise when more cycles were attempted</p> | ||
| + | |||
| + | <p> </p> | ||
| + | |||
| + | <p><strong>changing surfatin</strong></p> | ||
| + | |||
| + | <p>The 5th A domain in the NRPS for surfactin was attempted changed from a aspartate to a asparagine. This was done by following the OGRE protocol using a oligo that would change the specificity from asp to asn.</p> | ||
| + | |||
| + | <p> </p> | ||
</div> | </div> | ||
| Line 244: | Line 276: | ||
</div> | </div> | ||
<ul class="timeline"> | <ul class="timeline"> | ||
| + | |||
| + | <li> | ||
| + | <div class="timeline-badge"><i class="glyphicon glyphicon-check"></i></div> | ||
| + | <div class="timeline-panel"> | ||
| + | <div class="timeline-heading"> | ||
| + | <h4 class="timeline-title">Biology of the Future </h4> | ||
| + | <p><small class="text-muted"><i class="glyphicon glyphicon-time"></i> 2015-09-14</small></p> | ||
| + | </div> | ||
| + | <div class="timeline-body"> | ||
| + | <p style="text-align: justify;"><img alt="" src="None" style="height: 158px; width: 250px; float: left;" />Our team member Pernille spent an afternoon giving a presentation on cell factories and synthetic biology to 30 highschool students. She says: "Synthetic biology is all about being creative and use your imagination to design bio-solutions that can impact the world. For that exact reason, it is neccesary to discuss which ethical values tat should drive research". The highschool students had lots of questions and opinion for the discussion about ethics.</p> | ||
| + | |||
| + | <p style="text-align: justify;"> </p> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </li> | ||
| + | |||
| + | <li class="timeline-inverted"> | ||
| + | <div class="timeline-badge"><i class="glyphicon glyphicon-check"></i></div> | ||
| + | <div class="timeline-panel"> | ||
| + | <div class="timeline-heading"> | ||
| + | <h4 class="timeline-title">VWR to the rescue! </h4> | ||
| + | <p><small class="text-muted"><i class="glyphicon glyphicon-time"></i> 2015-09-01</small></p> | ||
| + | </div> | ||
| + | <div class="timeline-body"> | ||
| + | <p>We are happy to announce that the VWR is contributing our team with their awesome products!</p> | ||
| + | |||
| + | <p><img alt="" src="None" style="width: 400px; height: 533px;" /></p> | ||
| + | |||
| + | <p> </p> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </li> | ||
<li> | <li> | ||
| Line 560: | Line 626: | ||
</div> | </div> | ||
| − | < | + | <div class="container"> |
<div class="row"> | <div class="row"> | ||
<div class="col-md-2 col-md-offset-2"> | <div class="col-md-2 col-md-offset-2"> | ||
| Line 584: | Line 650: | ||
</div> | </div> | ||
</div> | </div> | ||
| − | </ | + | </div> |
<div class="sponsors"> | <div class="sponsors"> | ||
| − | <div class="container"> | + | <div class="container-fluid"> |
<div class="row"> | <div class="row"> | ||
Revision as of 14:21, 16 September 2015
Summary
For OGRE (Oligo Recombineering) to be effective there is two genetic improvements that can be done. The first and most important is that the organism need to express a recombinase protein such as the beta protein derived from the lambda RED E. coli phage. The second improvement is to destroy/inhibit the mismatch repair system [1]. In B. subtilis the genes mutS and mutL are taking care of the mismatch repair [2]. For proof of concept purposes deleting the mismatch repair system will be adequate. According to (REF) the protein encoded by mutL will not be functional without a functional mutS, this is why deletion of only mutS will be sufficient to take out the mismatch repair system. With respect to recombinase we will test two different recombinases: lambda-beta (codon optimized for B. subtilis 168) and gp35 (a recombinase from the B. subtilis phage SPP1) [3]. For duing proof of concept of OGRE in B. subtilis four different strains was producere via genetic recombineering. All derived from Bacillus subtilis 168:
- ∆amyE::beta-neoR
- ∆amyE::GP35-neoR
- ∆mutS::beta-neoR
- ∆mutS::GP35-neoR
The four different strains where compared to each other to get compare there transformation efficienc in MAGE.
Methods
making beta and GP35
The beta CDS was codon optimized with JCat() and ordered from COMPANY. GP35 sequence was found from (GP35 article). The sequence was ordered from the same company as beta.
incertion of recombinases in the amyE locus of B. subtilis
Beta and GP35 was inserted into the plasmid known as pDG268neo by using PCR with G5 HF 2x master mix from NEB (primer sequence hyperlink?). The plasmid was transformed into E coli DH5alpha by heat chock and incubated over night. The plasmid was purified from E coli by using QiAPrep mini prep kit. Digesting with restriction enzymes validated the insertion. This plasmid integrates the CDS into the amyE locus in Bacillus subtilis. pDG268neo with ether beta or GP35 was linearized and transformed into B subtilis W168 using the CFrench protocol (make CFrench to hyperlink)
chromosomal DNA from B. subtilis 168 was isolated (protocol)
knock out mutS
The regions upstream and downstream of the mutS locus were amplified by PCR from the chromosomal DNA with primers that had overlap with GP35 and beta.
GP35 and beta were PCR amplified with primers that had overhangs that were homologies to the mutS locus in bacillus subtilis. GP35 and beta was assembled into the pSB1C3 plasmid by using gibson assembly and transformed into E coli and incubated over night. The plasmid was isolated by using mini prep. The plasmids were verified using restriction enzymes.
beta and GP35 in pSB1C3 was linearized using XhoI, This also removed the Chloramphenicol gene from the plasmid. Beta and GP35 was transformed into B. subtilis.
Genomic DNA extraction (B. brevis)
Ligation of DNA
PCR -
- PCR
- Colony PCR
- Touch down
- …
Plasmid Purification Miniprep
Restriction enzyme digest (NEB)
Tyrocidine construct
Transformation and Preparation of E.coli
-Preparation of Competent Cells
-Transformation of Chemically Competent E.coli
Transformation of Bacillus
Optiminsing OGRE in Bacillus
Different amount of oligo was tested to optimize OGRE method for Bacillus subtilis.
The mismatch insertion frequency and oligo length was tested. The standard OGRE protocol was follow with the mismatches verified from 1-6 mismatches and oligoes from 50-100nt.
OGRE with 4 cycles were attempted every cycle to approximately 6 hours. This was done to test if the transformation efficiency would rise when more cycles were attempted
changing surfatin
The 5th A domain in the NRPS for surfactin was attempted changed from a aspartate to a asparagine. This was done by following the OGRE protocol using a oligo that would change the specificity from asp to asn.
Timeline
-
Biology of the Future
2015-09-14
Our team member Pernille spent an afternoon giving a presentation on cell factories and synthetic biology to 30 highschool students. She says: "Synthetic biology is all about being creative and use your imagination to design bio-solutions that can impact the world. For that exact reason, it is neccesary to discuss which ethical values tat should drive research". The highschool students had lots of questions and opinion for the discussion about ethics.
-
VWR to the rescue!
2015-09-01
We are happy to announce that the VWR is contributing our team with their awesome products!
-
Visit from Fisher Scientific!
2015-08-26
Today we had a visit by Susanne Basse from Fisher Scientific. She came by our office with a wagon full of items, Thanks!

-
-
-
-
-
Canoeing
2015-08-07
Today we took a day off from lab to go canoeing, followed by a barbeque at Chris' place.

-
-
Wiki Wizard
2015-07-30
We have uploaded the first beta version of the Wiki Wizard to GitHub
-
First BioBrick cloned into B. subtilis
2015-07-30
An expression cassette with lambda beta recombinase was sucesfully transformed into Bacillus subtilis. First step towards establishing MAGE in Bacillus subtilis completed!
-
BioBrick High School Project
2015-07-23
3 high school students; Simran, Noor, and Charlotte, joined us and Biotech Academy for 5 days to make biosensors that changes color, when water is poluted. Read more at ADD and http://biotechacademy.dk.

-
First sequencing results
2015-07-22
We received our first sequencing results. One step closer to our first BioBrick!
-
Brevibacillus parabrevis arrives
2015-07-16
Brevibacillus parabrevis produces a lot of different antibiotics including tyrocidine. We will transfer the tyrocidine operon to our chassis Bacillus subtilis.
-
-
-
-
We get our strain
2015-04-28

We get a plate of our chassis strain Bacillus subtilis W168.
We can now begin the work in the laboratory. -
BioBrick Workshop
2015-04-24
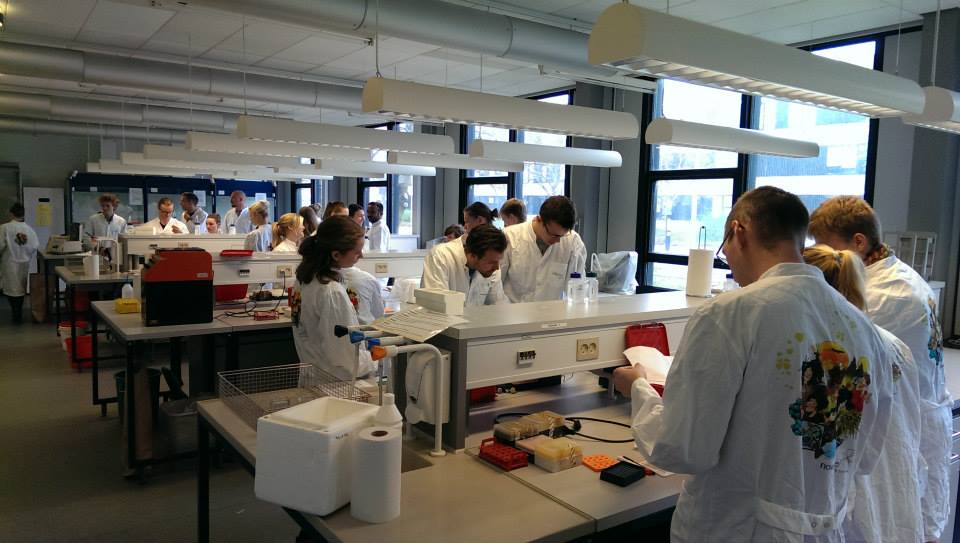

Southern University of Denmark and University of Copenhagen joined us for a three-day BioBrick tutorial, where we spent three days in the laboratory cloning, learning about iGEM, wiki design, and getting to know each other through social activities.
-
-
First official team meeting
2015-01-14
Today we had our first official team meeting. We talked about the project and got to know each other.
-
Introductory meeting
2014-11-25
This day, last years team held an introductory session about iGEM for interested people.
Department of Systems Biology
Søltofts Plads 221
2800 Kgs. Lyngby
Denmark
P: +45 45 25 25 25
M: dtu-igem-2015@googlegroups.com