Team:Czech Republic/Interlab study
Interlab study
Contents
Background
“All iGEM teams are invited and encouraged to participate in the Second International InterLab Measurement Study in synthetic biology.” Our team accepted this remarkable challenge that should show differences in cloning and measurement among teams all over the world. The main task was to characterize expression profiles of three promoters with different strength. In our case, we measured fluorescence using flow cytometery.
Design
Cloning of a GFP gen into the vector containing promoter allows us to measure strength of promoter via fluorescence. For construction of such device we chose to use BioBrick Standard Assembly. The suffix of promoters was cut with the enzymes SpeI and PstI. The prefix of the GFP gen was cut with the XbaI enzyme and the suffix with the PtI enzyme. Enzymes SpeI and XbaI have compatible sticky ends. This allows us to connect two parts of the final device together through ligation.
Materials and methods
Used plasmids
- Device 1 (20K+21J): J23101 + I13504, in the pSB1C3 backbone
- Device 2 (22A+21J): J23106 + I13504, in the pSB1C3 backbone
- Device 3 (22K+21J): J23117 + I13504, in the pSB1C3 backbone
- Positive control (C+): I20270, in the pSB1C3 backbone
- Negative control (C-): K823008, in the pSB1C3 backbone
Used strain
E. coli E5alpha
Used material
- LB-M agar plates with chloramphenicol
- LB-M agar plates with ampicillin
- 1.5 ml eppendorf tubes
- 0.5 ml PCR tubes
- 50 ml centrifuge conical base and rim tubes
- NucleoSpin Plasmid DNA, RNA, and protein purification Kit
- NucleoSpin Gel and PCR Clean-up Kit
- LB-M medium with chloramphenicol
- NaOH agarose gel and buffer
- Sphero Rainbow Calibration Particles, 8 Peaks, 3.0-3.4
Used methods
- Transformation
- Miniprep
- Restriction digest
- Ligation
- NucleoSpin Gel Clean-up
- NucleoSpin Plasmid DNA purification
- Flow cytometry
All used protocols can be found here: Protocols
Used machines
- Incubator: Binder, ATP.line CB(E6), CO2 – Incubator with O2 control, model CB160, 230V
- Orbita Shaker PSU-10i, 280 rpm, 360deg
- Flow Cytometr C6
- Thermo-Shaker with cooling for microtubes and PCR plates TS-100C
- Centrifuge 5424/5424 R
Used software
- CFlow Plus
- Microsoft Excel
- Sphero PMT QC Template
Description
Plasmids containing promoters and GFP were taken from The 2015 DNA Distribution Kit and transformed into E. coli. Colonies were streaked for patches and amplified plasmids were obtained using NucleoSpin Plasmid DNA purification. Restriction digest followed using XbaI and PstI enzymes for digest of plasmid with GFP and SpeI and PstI ezymes for digest of plasmids containing promoters. Desired parts were then cut out of gel and purified using NucleoSpin Gel Clean-up. Ligation of Device 1 and 2 followed our standard protocol. However ligation of Device 3 had to be repeated several times and the protocol had to be modified (2 hours ligation, 5:1 insert:vector ratio). All devices were transformed into E.coli. Fluorescence of colonies was checked up under UV light. Plasmids were then obtained by NucleoSpin Plasmid DNA purification. Devices were verified on gel after restriction digest using SpeI and EcoRI enzymes and by sequencing.
All verified devices were again transformed into E. coli accompanied by positive and negative control. 5 ml of liquid LB-M medium with chloramphenicol were inoculated with three chosen colonies of each device. There were no biological replicates for positive and negative control. Liquid cultures were incubated for 17 hours in Orbita Shaker PSU-10i placed in incubator. OD of these cultures was measured and diluted to 0.1. Fluorescence of biological and also technical replicates was measured using Flow Cytometr C6 following our flow cytometry protocol.
Processing of data
Raw data obtained from Flow Cytometr C6 were transformed into MEFL (Molecules of Equivalent Fluorescein) unites. We used Sphero Rainbow Calibration Particles, 8 Peaks, 3.0-3.4 to obtain a Calibration Graph using Sphero PMT QC Template. MEFL data were then processed in Microsoft Excel.
Protocol [Spherotech]
- Run the 3.0 μm RCPs, which the MEFL are known. Adjust the signal settings such as laser power to place all peaks within the 4-decade log scale. Record the fluorescence Mean Channel Number for each peak.
- Plot the assigned MEFL value for each peak vs the Mean Channel Number on the SPHERO PMT QC Teplate to obtain a Calibration Graph. (see appendix)
- Run the unknown sample using the same instrument settings (see Flow Cytometr protocol). Record the Mean Channel Number of sample.
- Calculate the MEFL of the unknown by crosscalibrating its Mean Channel Number against the Calibration Graph of the RCPs.
Results
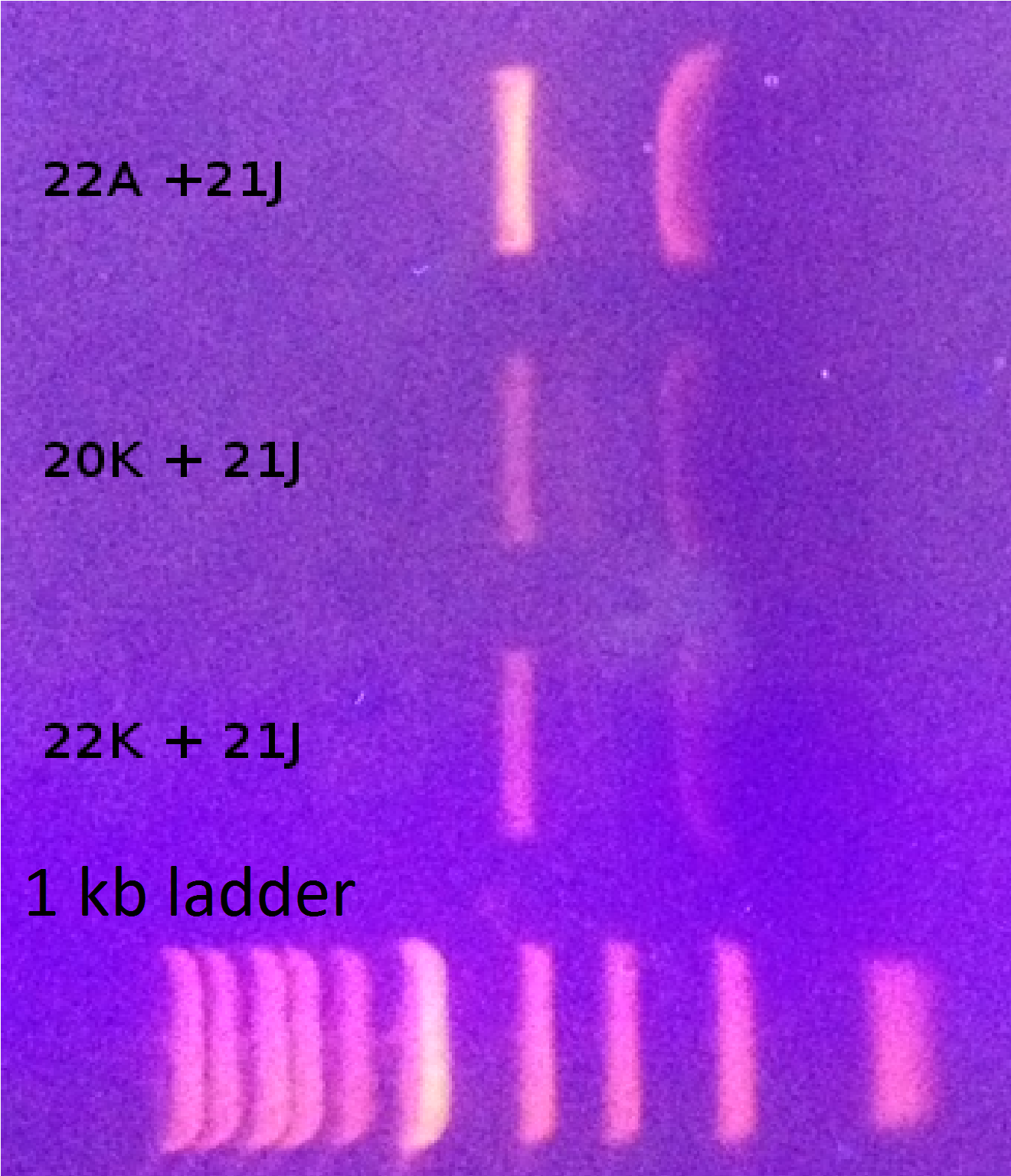
Gel verification
Sequencing
Device 1 (20K+21J)
18354158.seq - ID: 04IG33- on 2015/7/28-19:47:38 automatically edited with PhredPhrap, start with base no.: 6 Internal Params: Windowsize: 20, Goodqual: 19, Badqual: 10, Minseqlength: 50, nbadelimit: 1 TgattacTaTAAaaTaGGCGTatcacGaggCAGAaTTtCAGATAAAAAAAaTCCTTAGCTTTCGCTAAGGATGATTTCTGGAATTCGCGGCCGCTTCTAGAGTTTACAGCTAGCTCAGTCCTAGGTATTATGCTAGCTACTAGAGAAAGAGGAGAAATACTAGATGCGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCAGTGGAGAGGGTGAAGGTGATGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGAAAACTACCTGTTCCATGGCCAACACTTGTCACTACTTTCGGTTATGGTGTTCAATGCTTTGCGAGATACCCAGATCATATGAAACAGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGTACAGGAAAGAACTATATTTTTCAAAGATGACGGGAACTACAAGACACGTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGAAGATGGAAACATTCTTGGACACAAATTGGAATACAACTATAACTCACACAATGTATACATCATGGCAGACAAACAAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTGAAGATGGAAGCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTCCACACAATCTGCCCTTTCGAAAGATCCCAACGAAAnGAGAGACCACATGGTCCTTCTTGAGTTTGTAACAGCTGCTGGGATTACACATGGCATGGATGAACTATACAAATAATAATACTAGAGCCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTtATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATATACTAGTAGCGGCCGCTGCAGTCCGGCAAAAAaGGgCAAGgtgTCACCACCcTGCCcTTTTtCTTtAAAaCCGAAAanaTTACTTcncGTtntGCAGgCTtccncgCTCactGACTcgCTgnncTCGGTCGTTCGGCTGCGg
Device 2 (22A+21J)
18353363.seq - ID: 04IG49- on 2015/7/28-17:51:38 automatically edited with PhredPhrap, start with base no.: 7 Internal Params: Windowsize: 20, Goodqual: 19, Badqual: 10, Minseqlength: 50, nbadelimit: 1 CTgattacTaTAAAataGGCGTaTcaCgaGggCAGAaTTTCAGATAAAAAAAaTCCTTAGCTTTCGCTAAGGATGATTTCTGGAATTCGCGGCCGCTTCTAGAGTTTACGGctAGCTCAGTCCTAGGTATAGTGCTAGCTACTAGAGAAAGAGGAGAAATACTAGATGCGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCAGTGGAGAGGGTGAAGGTGATGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGAAAACTACCTGTTCCATGGCCAACACTTGTCACTACTTTCGGTTATGGTGTTCAATGCTTTGCGAGATACCCAGATCATATGAAACAGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGTACAGGAAAGAACTATATTTTTCAAAGATGACGGGAACTACAAGACACGTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGAAGATGGAAACATTCTTGGACACAAATTGGAATACAACTATAACTCACACAATGTATACATCATGGCAGACAAACAAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTGAAGATGGAAGCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTCCACACAATCTGCCCTTTCGAAAGATCCCAACGAAAnGAGAGACCACATGGTCCTTCTTGAGTTTGTAACAGCTGCTGGGATTACACATGGCATGGATGAACTATACAAATAATAATACTAGAGCCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATATACTAGTAGCGGCCGCTGCAGTCCGGCAAAAAaGGgCAAGGTGTCACCACCcTGCCcTTTTtCTTtAAaaCcgAAaaGATTACTTcnnnGTTATGCAGgCTTCCTCGCTCactGACTCGCTGCgCtcGgTCGTTCGgntncGg
Device 3 (22K+21J)
18593880.seq - ID: 04IF93- on 2015/8/10-20:57:38 automatically edited with PhredPhrap, start with base no.: 29 Internal Params: Windowsize: 20, Goodqual: 19, Badqual: 10, Minseqlength: 50, nbadelimit: 1 gCGtatcAcgaGggCagAaTTtCAGATAAAAAAAaTCCTTAGCTTTCGctAAGGATGATTTCTGGAATTCGCGGCCGCTTCTAGAGTTGACAGCTAGCTCAGTCCTAGGGATTGTGCTAGCTACTAGAGAAAGAGGAGAAATACTAGATGCGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCAGTGGAGAGGGTGAAGGTGATGCAACATACGGAAAACTTACCCTTAAATTTATTTGCACTACTGGAAAACTACCTGTTCCATGGCCAACACTTGTCACTACTTTCGGTTATGGTGTTCAATGCTTTGCGAGATACCCAGATCATATGAAACAGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGTACAGGAAAGAACTATATTTTTCAAAGATGACGGGAACTACAAGACACGTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATAGAATCGAGTTAAAAGGTATTGATTTTAAAGAAGATGGAAACATTCTTGGACACAAATTGGAATACAACTATAACTCACACAATGTATACATCATGGCAGACAAACAAAAGAATGGAATCAAAGTTAACTTCAAAATTAGACACAACATTGAAGATGGAAGCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTCCACACAATCTGCCCTTTCGAAAGATCCCAACGAAAnGAGAGACCACATGGTCCTTCTTGAGTTTGTAACAGCTGCTGGGATTACACATGGCATGGATGAACTATACAAATAATAATACTAGAGCCAGGCATCAAATAAAACGAAaGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGgCCTTtCTGCGTTtATATACTAGTAGCGgCCGCTGCAGTCCGGCAAAAAnGGgCAAGGTGTCAccaCCcTGCCcTTTTtCTTtAAAaCCGAAAnGaTTACTTcgCGTTATGCAGGCTTccTCGCTCACTGACTCGCTGcgCtcGgTCGTtCGGCTGCGgnnagcGGtaTCAgct
Data
| Device | Colony | MEFL | Mean of technical replicates | SD Colony | Mean of device | SD Device | |
|---|---|---|---|---|---|---|---|
| 20K +21J (Device 1) | Colony 1 | I. | 639549 | 657818 | 15504 | 664409 | 29185 |
| II. | 677453 | ||||||
| III. | 656453 | ||||||
| Colony 2 | I. | 689717 | 702990 | 10067 | |||
| II. | 714084 | ||||||
| III. | 705169 | ||||||
| Colony 3 | I. | 620331 | 632419 | 15053 | |||
| II. | 623286 | ||||||
| III. | 653639 | ||||||
| 22A+21J (Device 2) | Colony 1 | I. | 481600 | 464304 | 12795 | 392603 | 50712 |
| II. | 460265 | ||||||
| III. | 451048 | ||||||
| Colony 2 | I. | 346283 | 358070 | 9696 | |||
| II. | 357896 | ||||||
| III. | 370032 | ||||||
| Colony 3 | I. | 354544 | 355433 | 4688 | |||
| II. | 350188 | ||||||
| III. | 361568 | ||||||
| 22K+21J (Device 3) | Colony 1 | I. | 2635 | 2872 | 220 | 2229 | 454 |
| II. | 3165 | ||||||
| III. | 2814 | ||||||
| Colony 2 | I. | 1786 | 1891 | 74 | |||
| II. | 1933 | ||||||
| III. | 1953 | ||||||
| Colony 3 | I. | 1665 | 1926 | 204 | |||
| II. | 1948 | ||||||
| III. | 2163 | ||||||
| Positive control | Colony 1 | I. | 202361 | 196401 | 5853 | 196401 | / |
| II. | 188446 | ||||||
| III. | 198395 | ||||||
| Negative control | Colony 1 | I. | 224 | 213 | 8 | 213 | / |
| II. | 204 | ||||||
| III. | 211 | ||||||
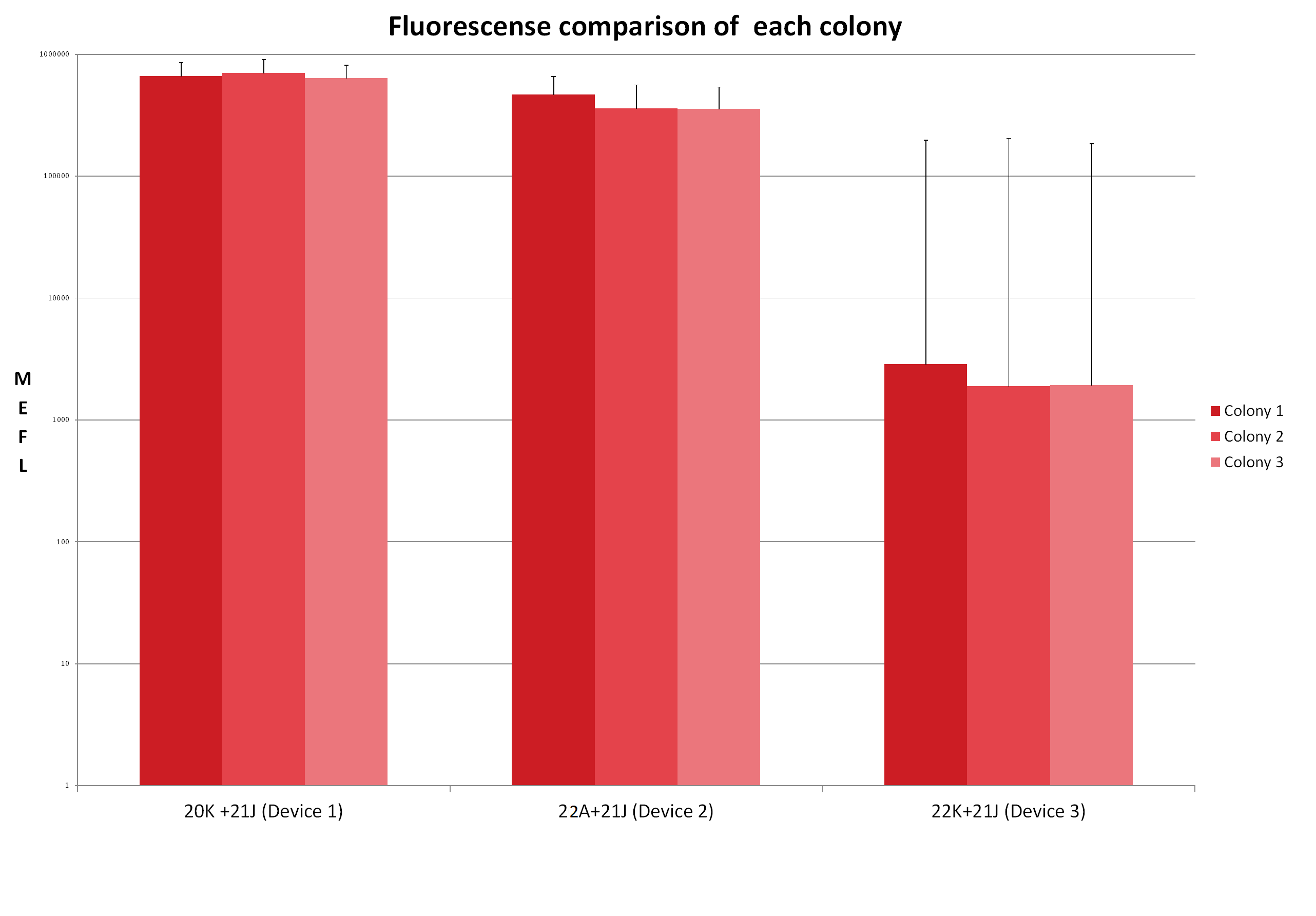
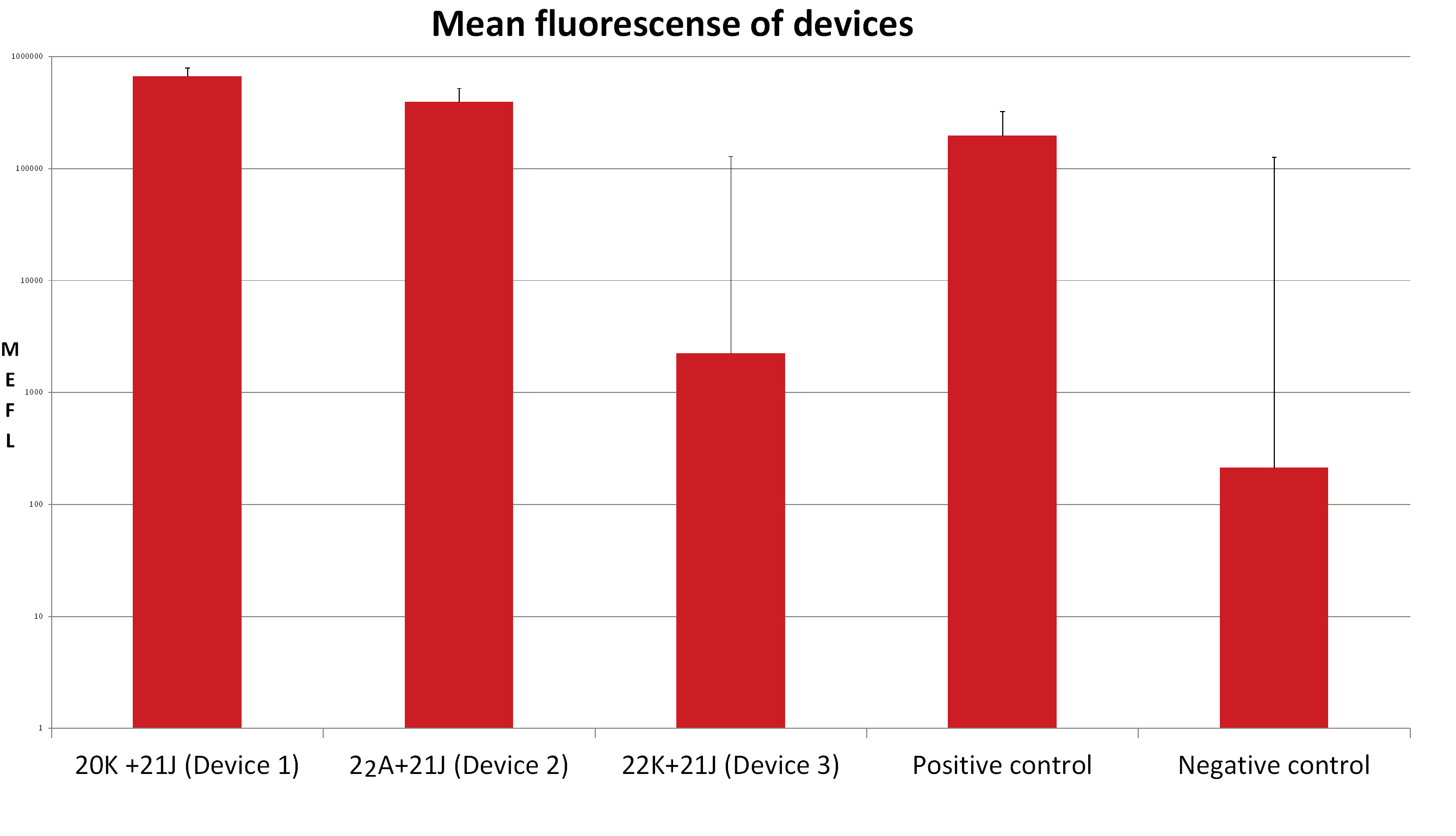
Visualization
Discussion
It is noticeable that the promoter of the Device 1 is strongest followed by the promoter of the Device 3. The device 1 has the weakest promoter.
Reference
- ↑ Spherotech, Inc. SHERO Technical Note, Measuring molecules of equivalent fluorochrome (MEF) using Sphero Rainbow and Ultrarainbow Calibration Particles[Online] [Cited: 9 4, 2015.]
Appendix
Individuals responsible for conducting InterLab study
- Anna Sosnová – Leadership, Device construction, Measurements, Data processing, Wiki content
- Václav Pelíšek – Experimental assistance
- Bc. Hynek Kasl – Experimental assistance
- Jan Bejvl – Data processing assistance
- Ing. Pavel Zach – Experimental advisor
- Mgr. Kateřina Pěchotová – Technical advisor
- M.Sc. Daniel Georgiev, PhD – support