Difference between revisions of "Team:Czech Republic/Project/Synthetic haploids"
| Line 1: | Line 1: | ||
{{:Team:Czech_Republic/Template:Top|Signal transduction}} | {{:Team:Czech_Republic/Template:Top|Signal transduction}} | ||
| − | |||
| − | |||
<html> | <html> | ||
<div class="left"> | <div class="left"> | ||
</html> | </html> | ||
| − | |||
= Overview = | = Overview = | ||
| − | |||
<html> | <html> | ||
</div> | </div> | ||
<div class="right"> | <div class="right"> | ||
</html> | </html> | ||
| − | |||
= Key Achievements = | = Key Achievements = | ||
| − | |||
Key Achievement #1 | Key Achievement #1 | ||
Key Achievement #2 | Key Achievement #2 | ||
| − | |||
<html> | <html> | ||
</div> | </div> | ||
</html> | </html> | ||
| − | |||
== Introduction: Yeast pheromone pathway== | == Introduction: Yeast pheromone pathway== | ||
[[File:Czech Republic Hapdip.png|600px|thumb|right|text]] | [[File:Czech Republic Hapdip.png|600px|thumb|right|text]] | ||
Revision as of 16:19, 9 September 2015
Signal transduction
Overview
Key Achievements
Key Achievement #1 Key Achievement #2
Introduction: Yeast pheromone pathway
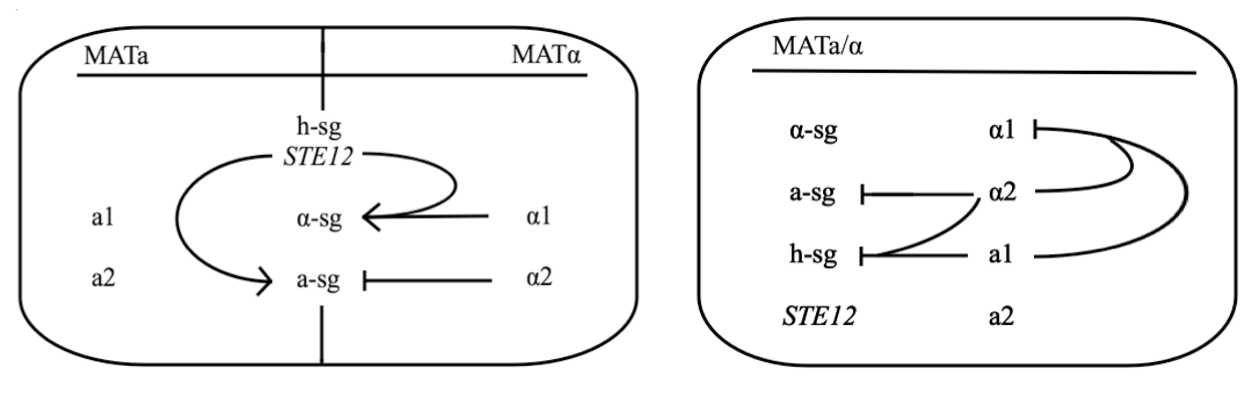
Yeast Saccharomyces cerevisiae exists either in haploid or diploid state. The two mating types are called MATa and MATα and differ only within approx. 2 kbp long region on chromosome III called MAT locus. MATa locus expresses transcription factors a1 and a2, whereas MATα locus expresses transcription factors α1 and α2. Both types express three groups of genes, which are:
- haploid-specific genes (h-sg)
- a-specific genes (a-sg)
- α-specific genes (α-sg)
As their names indicate, these genes are only active in haploids, MATa cells, or MATα cells, respectively.
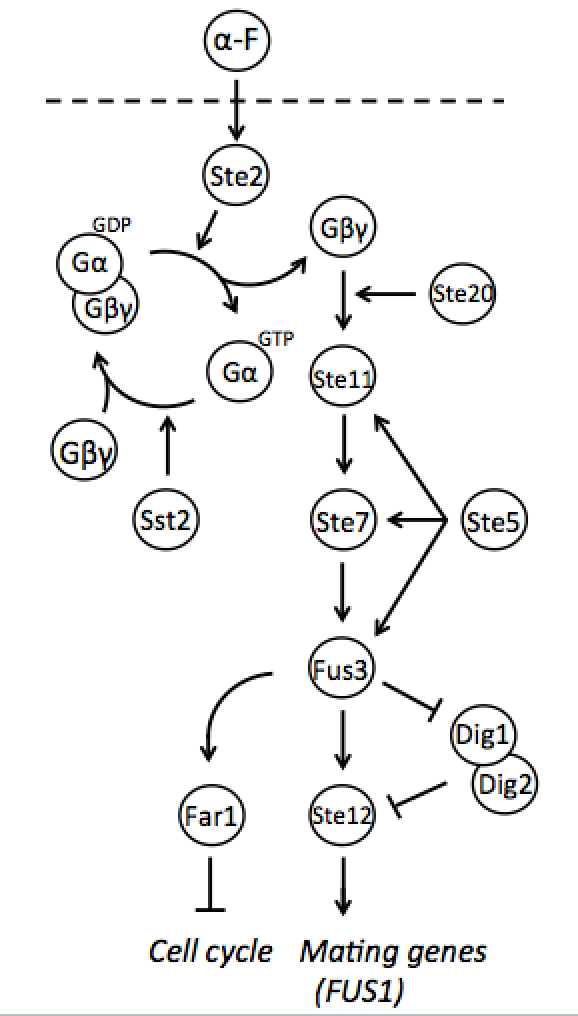
Both mating types constantly produce small amounts of mating type-specific pheromone and when the two cells of opposite types are in close proximity they identify each other by sensing each other’s pheromone. The response to detected pheromone is facilitated by so called pheromone pathway, which is a cascade of chemical reactions that results in the preparation for mating. Since only haploids mate, the yeast pheromone pathway is only functional in haploid cells.
Design: Reconstruction of yeast pheromone pathway in diploid cell
The schematic shows that h-sg genes are switched-off in diploid cell and since all genes included in yeast pheromone pathway signal transduction (except from the pheromone receptor)are h-sg, the pathway can be reconstructed by switching them on. It can be managed by interrupting one of the h-sg regulators - a1 or α2. While a1 has no function in haploid, its deletion has no effect in haploid cell, but in diploid it interrupts the repression of h-sg.
Naturally in haploids, after exposure to mating pheromone yeast pheromone pathway results in induction of expression of mating genes that initiate the mating process. Since it is not desired for the synthetic diploid to mate, the initiation of mating process needs to be disrupted. It can be achieved by repressing Ste12 transcription in the diploid. While Ste12 is fundamental in both haploid types in the process of mating, the designed mechanism must preserve transcription in haploid cells and repress it in diploid.
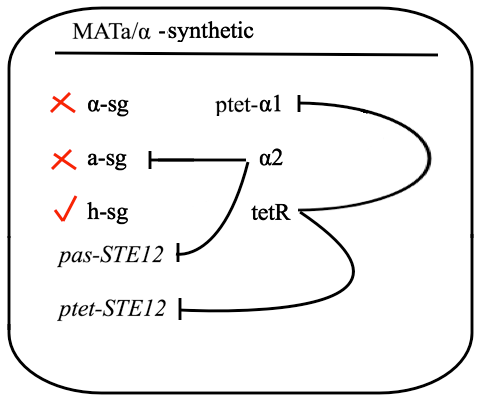
For this purpose, mechanism of tetracycline-controlled transcriptional activation was used: a1 gene in MATa was replaced by TetR and STE12 was put under the control of synthetic a-specific promoter that is repressed in the same way as a-sg. In MATα, α1 repression was preserved by placing it under the control of pTet and STE12 was also put under the control of pTet. As a result, transcription of Ste12 is preserved in both haploids and repressed in diploid - pTet-STE12 thanks to TetR repressor that is expressed from MATa chromosome, and a-specific STE12 thanks to α2. Also h-sgare expressed because of disruptedrepression by deleting a1 preserving transcription of all the genes forming yeast pheromone pathway.
See following pages for detailed information about constructing the synthetic haploids:
Concept
DNA
Materials and methods
Used strains
Used material
- LB-M agar plates with chloramphenicol
- LB-M agar plates with ampicillin
- 1.5 ml eppendorf tubes
- 0.5 ml PCR tubes
- 50 ml centrifuge conical base and rim tubes
- NucleoSpin Plasmid DNA, RNA, and protein purification Kit
- NucleoSpin Gel and PCR Clean-up Kit
- LB-M medium with chloramphenicol
- NaOH agarose gel and buffer
- Sphero Rainbow Calibration Particles, 8 Peaks, 3.0-3.4
Used methods
- Transformation
- Miniprep
- Restriction digest
- Ligation
- NucleoSpin Gel Clean-up
- NucleoSpin Plasmid DNA purification
- Flow cytometry
All used protocols can be found here:
Protocols
Used software
- CFlow Plus
- Microsoft Excel
- Sphero PMT QC Template
Construction
Construction of reporter plasmids
The pADH1, pSTE2, pSTE5, and pFUS1 promoters were obtained by PCR from yeast genome (isolated according to standard protocol from 7283 MATx strain). The asCYC1 and pTv3 promoters were obtained by PCR from g-blocks. The primers used for this are as follows:
INSERT TABLE OF PRIMERS
All promoters were amplified in a single PCR run, with the following conditions:
| Property | Value |
|---|---|
| Polymerase | Q5 |
| Extension Time | 90s |
PCR products were then gel verified
INSERT PHOTO_1
and were then purified (insert purification kit name) and restricted by corresponding restriction enzymes, which are listed in the following table
INSERT TABLE OF RESTRICTION ENZYMES
The corresponding vector for reporter plasmids was prepared by restriction and ligation of yeGFP and CYC1 terminator into a pRS416 CEN plasmid.
Construction of INSERT MATa and INSERT MATx
The backbone pRSII406 was obtained from Addgene. MATa was ordered in the form of gblock yG_MATa with sequence
yG_MATa :
AATTCATCTAGAGAAGAAAGCAAAGCCTTAATTCCAAGGAAAAAGAAGAAGTTGCAAAGAAATGTGGCATTACTCCACTTCAAGTAAGAGTTTGGGTATGTAATATGAGAATCAAACTTAAATATATCCTATACGTAGTATGGCGGAAAACATAAACAGAACTCTGTTTAACATTCTAGGTACTGAGcaaattaaagccttcgagcgtcccaaaaccttctcaagcaaggttttcagtataatgttacatgcgtacacgcgtctgtacagaaaaaaaagaaaaatttgaaatataaataacgttcttaatactaacataactataaaaaaataaatagggacctagacttcaggttgtctaactccttccttttcggttagagcggatgtggggggagggcgtgaatgtaagcgtgacataactaatCTAAAATTCCCGGGATCCGCTGTACGCGGACCCACTTTCACATTTAAGTTGTTTTTCTAATCCGCATATGATCAATTCAAGGCCGAATAAGAAGGCTGGCTCTGCACCTTGGTGATCAAATAATTCGATAGCTTGTCGTAATAATGGCGGCATACTATCAGTAGTAGGTGTTTCCCTTTCTTCTTTAGCGACTTGATGCTCTTGATCTTCCAATACGCAACCTAAAGTAAAATGCCCCACAGCGCTGAGTGCATATAATGCATTCTCTAGTGAAAAACCTTGTTGGCATAAAAAGGCTAATTGATTTTCGAGAGTTTCATACTGTTTTTCTGTAGGCCGTGTACCTAAATGTACTTTTGCTCCATCGCGATGACTTAGTAAAGCACATCTAAAACTTTTAGCGTTATTGCGTAAAAAATCTTGCCAGCTTTCCCCTTCTAAAGGGCAAAAGTGAGTATGGTGCCTATCTAACATTTTAATAAGTTGATTGTATGCTTGGTATAGCTTGAAATATTGTGCAGAAAAAGAAACAAGGAAGAAAGGGAACGAGAACAATGACGAGGAAACAAAAGATTAATAATTGCAGGTCTATTTATACTTGATAGCAAGACAGCAAACTTTTTTTTATTTCAAATTCAAGTAACTGGAAGGAAGGCCGTATACCGTTGCTCATTAGAGAGTAGTGTGCGTGAATGAAGGAAGGAAAAAGTTTCGTGTGCTTCGAGATACCCCTCATCAGCTCTGGAACAACGACATCTGTTGGTGCTGTCTTTGTCGTTAATTTTTTCCTTTAGTGTCTTCCATCATTTTTTTGTCATTGCGGATATGGTGAGACAACAACGGGGGAGAGAGAAAAGAAAAAAAAAGAAAAGAAGTTGtaaacccacaccgggtgtcataatcaaccaatcgtaacttcatctcttccacccatgtctctttgagcaataaagccgataacaaaatctttgtcgctcttcgcaatgtcaacagtacccttagtatattctccagtagatagggagcccttgcatgacaattctgctaacatcaaaaggcctctaggttcctttgttacttcttctgccgcctgcttcaaaccgctaacaatacctggTccACTAGTCCCGGGAGCAAGATCAAGATGTTTTCACCGATCTTTCCGGTCTCTTTGGCCGGGGTTTACGGACGATGGCAGAAGACCAAAGCGCCAGTTCATTTGGCGAGCGTTGGTTGGTGGATCAAGCCCACGCGTAGGCAATCCTCGCAGATCTCGAACCATGTAATTTCCGAATACGGTAATTACACGCATCGAGCAGATCCGCCAGGCGTGTATATATAGCGTGGATGGCCAGGCAACTTTAGTGCTGACACATACAGGCATATATATATGTGTGCGACGACACATGATCATATGGCATGCATGTGCTCTGTATGTATATAAAACTCTTGTTTTCTTCTTTTCTCTAAATATTCTTTCCTTATACATTAGGACCTTTGCAGCATAAATTACTATACTTCTATAGACACACAAACACAAATACACACACTAAAaagctt
MATx was ordered in the form of two g-blocks, yG_MATx1 and yG_MATx2 with the following sequences
yG_MATx1 :
AAGCTTGGATTCTCACAATCCTGTCGGTCACTTCTCGGCTGTTCGCGTATATTTTTTGTTGATACTTTTACCGGTATTTTGTCTGTAATTTATTCTCTATCACTGATAGGGACTTCTCTATCACTGATAGGGAACCCAGCCTGATTTATACTATTAGGGATCGCAGGAAGGCGGTGGGAAGTCCGGGAGTCGCTGAGGGGAAGTGTCAGTGGTTTTGGGTATAAATGGCTGGTTGTTCCCTATCAGTAATAGAGAATTCCCTATCAGTGATAGAGACTGCGGATTTAGAAACTACCTGATAAAAGTATCAACAAAAATTGCGCATGCCGGCCTGGATTTTGCGCAAATTTACCTTAACGTCCCACAATATGTTTACTTCGAAGCCTGCTTTCAAAATTAAGAACAAAGCATCCAAATCATACAGAAACACAGCGGTTTCAAAAAAGCTGAAAGAAAAACGTCTAGCTGAGCATGTGAGGCCAAGCTGCTTCAATATTATTCGACCACTCAAGAAAGATATCCAGATTCCTGTTCCTTCCTCTCGATTTTTAAATAAAATCCAAATTCACAGGATAGCGTCTGGAAGTCAAAATACTCAGTTTCGACAGTTCAATAAGACATCTATAAAATCTTCAAAGAAATATTTAAACTCATTTATGGCTTTTAGAGCATATTACTCACAGTTTGGCTCCGGTGTAAAACAAAATGTCTTGTCTTCTCTGCTCGCTGAAGAATGGCACGCGGACAAAATGCAGCACGGAATATGGGACTACTTCGCGCAACAGTATAATTTTATAAACCCTGGTTTTGGTTTTGTAGAGTGGTTGACGAATAATTATGCTGAAGTACGTGGTGACGGATATTGGGAAGATGTGTTTGTACATTTGGCCTTATAGAGTGTGGTCGTGGCGGAGGTTGTTTATCTTTCGAGTACTGAATGTTGTCAGTATAGCTATCCTATTTGAAACTCCCCATCGTCTTGCTGCAG
yG_MATx2 :
CTGCAGAGTAGTGTCTGAGGTACAAACATCTTAGTAGTGTCGAGAGGGTTGATTGTTTATGTATTTTTGCGAAATATATATATATATATTCTACACAGATATATACATATTTGTTTTTCGGGCTCATTCTTTCTTCTTTGCCAGAGGCTCACCGCTCAAGAGGTCCGCTAATTCTGGAGCGATTGTTATTGTTTTTTCTTTTCTTCTTCTATTCGAAACCCAGTTTTTGATTTGAATGCGAGATAAACTGGTATTCTTCATTAGATTCTCTAGGCCCTTGGTATCTAGATATGGGTTCTCGATGTTCTTTGCAAACCAACTTTCTAGTATTCGGACATTTTCTTTTGTAAACCGGTGTCCTCTGTAAGGTTTAGTACTTTTGTTTATCATATCTTGAGTTACCACATTAAATACCAACCCATCCGCCGATTTATTTTTCTGTGTAAGTTGATAATTACTTCTATCGTTTTCTATGCTGCGCATTTCTTTGAGTAATACAGTAATGGTAGTAGTGAGTTGAGATGTTGTTTGCAACAACTTCTTCTCCTCATCACTAATCTTACGGTTTTTGTTGGCCCTAGATAAGAATCCTAATATATCCCTTAATTCAACTTCTTCTTCTGTTGTTACACTCTCTGGTAACTTAGGTAAATTACAGCAAATAGAAAAGAGCTTTTTATTTATGTCTAGTATGCTGGATTTAAACTCATCTGTGATTTGTGGATTTAAAAGGTCTTTAATGGGTATTTTATTCATTTTTTCTTAGTGTGTGTATTTGTATTTGCGTGTCTATAGAAGTATAGTAATTTATGCTGCAAAGGTCCTAATGTATAAGGAAAAAAAATTTAGAGAAAAAAAGAAAAAAAGAGTTTTATATACATACAGAGCACATACATGCCATATAATCATGTATATACGCGCACATATATATATGCCTGTATGTGTCAGCACTAAATTTACCTGAACATACGCGCTATATATACGCGCCTCGCGTATATGCTCGAGGATTCCCTACGCGTGGGCTTTTTTTACTAACCAACGCGCGCGAAATActagt
The g-blocks were restricted by the following enzymes, along with the vector
insert table 1
Restriction were purified (insert purification kit name) and ligated together using the standard ligation protocol.
STE12 was PCR amplified from yeast genome (isolated from 7283 MATx strain) using the following primers :
Forward : CTTGTAAAGCTTCCAAGGATGAAAGTCCAAATAACCAATAGTAGAACA Reverse : ACTGCACTCGAGAGATTTGTTACATTTATTACCTTTTTTTCTTGCTTT
The conditions for the PCR were the following
Results
Final constructs
Herein we should describe the final constructs we have obtained, and add verification gels etc.
Test of mating types
Appendix
Personnel
Tereza Puchrová - The leading scientist
Hynek Kasl - The helping hand
Anna Sosnová - The actual worker
Václav Pelíšek - The parafilmer of plates
Useful Links
Protocols page:
References
- ↑ Lin, C.-H., Choi, a., & Bennett, R. J. (2011). Defining pheromone-receptor signaling in Candida albicans and related asexual Candida species. Molecular Biology of the Cell, 22(24), 4918–4930. doi:10.1091/mbc.E11-09-0749