Difference between revisions of "Team:Czech Republic/Project/Location tags"
m |
(→References) |
||
| Line 158: | Line 158: | ||
| − | + | # {{:Team:Czech_Republic/Template:Reference|(Zhao et al, 2001)|Zhao, H.,Shen, Z.-M., Kahn, P. C. and Lipke, P. N. (2001). Interaction of α-Agglutinin and a-Agglutinin, Saccharomyces cerevisiae Sexual Cell Adhesion Molecules. Journal of Bacteriology, 183(9), 2874–2880. doi:10.1128/JB.183.9.2874-2880.2001}} | |
| − | + | # {{:Team:Czech_Republic/Template:Reference|(Lipke and Kurjan, 1992)|Lipke, P. N. and Kurjan, J. (1992). Sexual agglutination in budding yeasts: structure, function, and regulation of adhesion glycoproteins. Microbiology and Molecular Biology Reviews, 56(1), 180-194.}} | |
| − | + | # {{:Team:Czech_Republic/Template:Reference|(Boder and Wittrup, 1997)|Boder, E. T. and Wittrup, K. D. (1997). Yeast surface display for screening combinatorial polypeptide libraries. Nature Biotechnology, 15, 553-557. doi:10.1038/nbt0697-553}} | |
| − | + | # {{:Team:Czech_Republic/Template:Reference|(Baird et al, 2012)|Baird, C. et al. (2012). Yeast Display scFv Antibody Library User’s Manual. Richland, WA: Pacific Northwest National Laboratory.}} | |
| − | + | # {{:Team:Czech_Republic/Template:Reference|(Wasilenko et al, 2010)|Wasilenko, J. L. et al. (2010). Cell Surface Display of Highly Pathogenic Avian Influenza Virus Hemagglutinin on the Surface of Pichia Pastoris Cells Using alpha-Agglutinin for Production of Oral Vaccines. Biotechnology Progress, 26(2), 542-547. doi:10.1002/btpr.343.}} | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
{{:Team:Czech_Republic/Template:Bottom}} | {{:Team:Czech_Republic/Template:Bottom}} | ||
Revision as of 11:26, 16 September 2015
Location Tags
Contents
Abstract
To guarantee the ability of our system to recognise, bind, and respond to the presence of a tumour cell, we selected five antibody fragments to display on our yeast cells using the yeast display technology and monitor the cells' binding. Their expression was detected and monitored on a microfluidic chip with the use of antibody staining and fluorescence microscopy.
Key Achievements
Expressed streptavidin, EpCAM, Anti-EpCAM, c-myc and HuA receptors on the surface of yeasts. Monitored the dynamic binding of our receptors and corresponding markers. Demonstrated the ability of our receptors to bind chosen markers.
Introduction
Agglutinins
alpha-agglutinin and a-agglutinin are cell wall glycoproteins of S. cerevisiae that possess complementary surfaces and mediate cell-to-cell adhesion during mating [(Lipke and Kurjan, 1992)]. a-agglutinin consists of two subunits, of which Aga1p is anchored to the membrane by a glycosylphosphatidylinositol anchor (GPI) and to which the second subunit, Aga2p, is bound by two disulphide bonds. alpha-agglutinin is composed of only one subunit and is connected to a modified GPI anchor. They are both highly glycosylated and constitutively expressed in low levels [(Zhao et al, 2001)].
Yeast Display
The Yeast Surface Display was first described in 1997 by E. Border and K. Wittrup as a method for engineering cell surface and secretion proteins and most ultimately for screening polypeptide libraries for high-affinity antibodies [(Boder and Wittrup, 1997)]. The gene for the protein of interest is fused in frame to the AGA2 gene on a plasmid. The second part of the display, AGA1 gene is integrated into a chromosome in a special S. cerevisiae strain EBY100. Both of the constructs are originally under GAL1 promoter, thus requiring galactose to be present in the medium for the system to function. Moreover, two epitope tags are usually added to the protein of interest for better visualisation and manipulation - one upstream (hemagglutinin) and one downstream (c-myc) [(Baird et al, 2012)].
Design
We obtained the original yeast display system from professor Sheldon Park, which we are thankful for. This particular PYD plasmid used the BamHI and NheI sites for insertion of scFv sequences and possessed a hemagglutinin tag upstream of the scFv sequence insertion site and a FLAG epitope at the downstream end.
Also, we made several modification to the yeast display method in order for it to better suit our needs. The goal was to simplify it and reduce the amount of new DNA that needs to be introduced into a yeast cell. The first of two alternatives was based on creating a general cassette containing a promoter, a terminator, and necessary cloning sites. The second alternative meant attaching the specific antibody with a terminal epitope tag directly to Aga1 protein.
Modified Yeast Display
Expression Cassette
We wanted to build five variations of the pRSII col plasmid based on which final cassette will act as the insert. The design of our cassette included using the constitutive pTv3 promoter, and a terminator sequence. One of the five final cassettes would contain the sequence coding for the Aga1 protein, and four of the cassette inserts would contain sequences coding for Aga2 proteins fused with a single chain antibody of choice and. By carefully designed consequent PCR reactions and restriction digests we aimed to obtain in vitro the 1) promoter, 2) terminator, 3) promoter and terminator, 4) promoter, ORF and terminator. Subsequent restriction digests and ligations would lead to a complete expression plasmid containing our desired inserts. See the Figure 3.2 for a graphical explanation.
Aga1p only
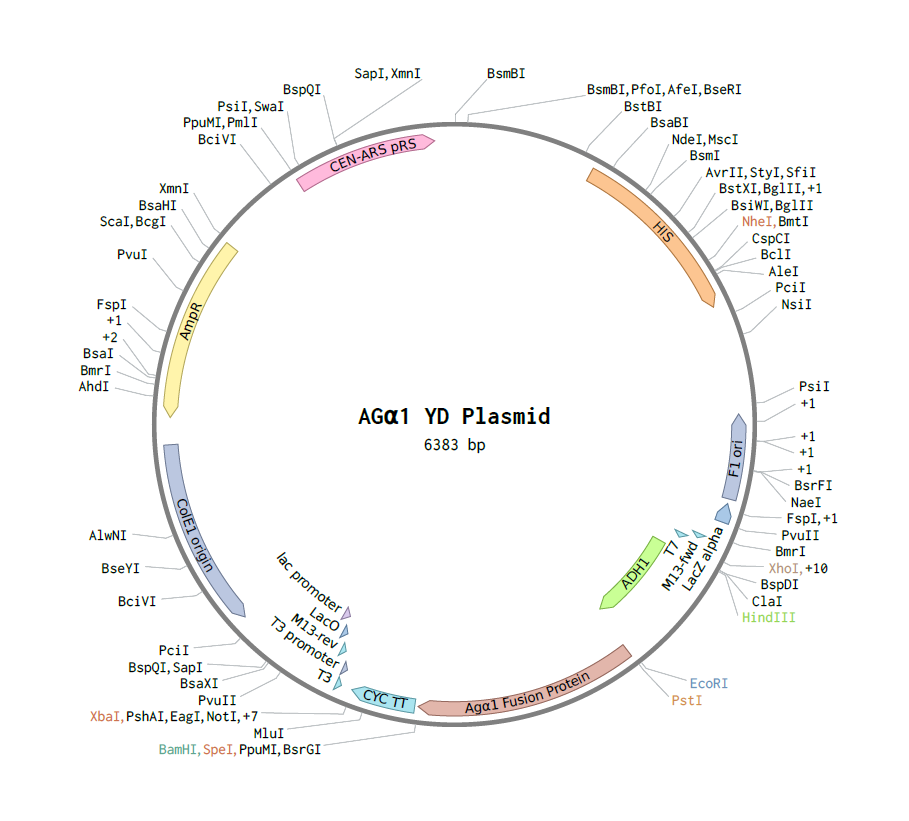
By fusing the antibody fragment directly to the Aga1 protein, we are able to eliminate the complications that arise because of the translation of several proteins in a row. The courage to pursue this alternative stemmed from the positive results obtained by J. Wasilenko and colleagues in their work of displaying the hemagglutinin of avian influenza virus on yeast cells using alpha-agglutinin. We aimed to incorporate into only one plasmid the sequence for an antibody fused with the C-terminal end of Agalpha1 (alpha-agglutinin) and a secretion signal. All of it to be controlled by one promoter [(Wasilenko et al, 2010)]. See Figure 3.3 for a schematic of the plasmid enabling yeasts to express an antibody on its surface using only the Aga1 protein.
Choice of Markers
Streptavidin
Streptavidin was our first choice of a marker because of its extremely strong bond with biotin. It could give us not only great results but also a benchmark affinity value to compare the other antibodies’ affinities to.
EpCAM
EpCAM, the transmembrane epithelial cell adhesion molecule, functions in cell signalling, proliferation, differentiation, and is present in many carcinomas. We selected it because of its significance in tumor growth promotion and its ability to bind Anti-EpCAm, which we also expressed on S. cerevisiae using yeast display.
c-myc
c-Myc is a transcription factor that regulates cell growth, apoptosis, and cellular differentiation. Its levels are often upregulated in different cancer types, where it plays a role in gene duplication and Ras interaction.
HuA
HuA stands for human antigen A of red blood cells. The decision to enable the expression of an antibody against antigen A on yeast surface allows us to test our system directly in blood and prove that the positive agglutination reaction is specific and clearly visible.
Construction
Construction of Traditional Yeast Display Plasmids
We obtained the sequences of streptavidin and individual antibodies from research papers online. After modifying the sequences to contain cloning sites needed for cloning into the YD vector and part submission, they were synthesized. Final sequences are pasted here:
Streptavidin
GTAAAACGACGGCCAGTGCTAGCGCTGAAGCTGGTATCACCGGCACCTGGTACAACCAGCTGGGTTCCACCTTCATCGTTACCGCTGGTGCTGACGGTGCTCTGACCGGTACCTACGAATCCGCTGTTGGTAACGCTGAAAGCCGCTACGTTCTGACCGGTCGTTACGACTCCGCTCCGGCTACCGACGGTTCCGGAACCGCTCTGGGTTGGACCGTTGCTTGGAAAAACAACTACCGTAACGCTCACTCCGCTACCACCTGGTCTGGCCAGTACGTTGGTGGTGCTGAAGCTCGTATCAACACCCAGTGGTTGTTGACCTCCGGCACCACCGAAGCCAACGCGTGGAAATCCACCCTGGTTGGTCACGACACCTTCACCAAAGTTAAACCGTCCGCTGCTTCTGGATCCGTCGTGACTGGGAAAAC
EpCAM
AAAAGTGCTAGCCAAGAAGAATGTGTATGTGAAAATTATAAACTAGCTGTCAACTGTTTTGTAAATAACAATAGACAGTGTCAATGTACTTCCGTTGGCGCTCAAAACACAGTTATCTGTTCTAAACTAGCCGCGAAGTGTCTTGTTATGAAGGCAGAGATGAATGGCTCCAAACTGGGTAGGCGTGCCAAGCCTGAGGGTGCCCTACAGAACAATGATGGGCTTTACGATCCAGATTGTGATGAATCCGGTTTATTCAAAGCCAAGCAGTGTAATGGTACTTCCATGTGCTGGTGTGTAAACACCGCTGGAGTGAGGAGAACCGATAAAGACACGGAAATCACCTGTAGCGAAAGAGTCAGGACTTACTGGATCATTATCGAACTGAAGCACAAAGCTAGGGAGAAGCCTTATGACAGCAAAAGTTTGAGGACTGCCTTACAGAAAGAAATCACAACTAGGTATCAATTAGACCCGAAATTTATAACGAGTATCCTTTACGAGAATAACGTCATTACCATTGATCTAGTGCAAAATTCTTCCCAGAAGACTCAAAATGACGTTGATATTGCAGATGTTGCCTATTACTTTGAAAAAGATGTGAAAGGAGAGAGTTTGTTCCACTCAAAAAAGATGGATCTGACAGTTAACGGGGAACAGTTAGATCTTGATCCGGGTCAAACGTTAATATATTATGTTGACGAAAAAGCACCAGAATTTAGTATGCAGGGATTGAAAGGATCCATAAAG
Anti-EpCAM (C215)
GAATTCGCGGCCGCTTCTAGATGCAGGTCAAGCTtCAGCAGTCAGGGGCTGAACTGGTGAGGCCTGGGGCTTCAGTGAAGCTGTCCTGCAAGGCTTCTGGCTACACCTTCACCAACTACTGGATAAACTGGGTGAAGCAGAGGCCTGGACAAGGCCTTGAGTGGATCGGAAATATTTATCCTTCTTATATTTATACTAACTACAATCAAGAGTTCAAGGACAAGGTCACATTGACTGTAGACGAATCCTCCAGCACAGCCTACATGCAGCTCAGCAGCCCGACATCTGAGGACTCTGCGGTCTATTACTGTACAAGATCCCCTTATGGTTACGACGAGTATGGTCTGGACTACTGGGGCCAAGGCACCACGGTCACCGTCTCCTCAGGTGGAGGCGGTTCAGGCGGAGGTGGCTCTGGCGGTGGCGGATCGGACATCGAGCTCACTCAGTCTCCATCCTCCCTGACTGTGACAGCAGGAGAGAAGGTCACTATGAACTGCAAGTCCAGTCAGAGTCTGTTAAACAGTAGAAATCAAAAGAACTACTTGACCTGGTACCAGCAGAAACCAGGGCAGCCTCCTAAACTGTTGATATACTGGGCATCCACTAGGGAATCTGGGGTCCCTGATCGCTTCACAGGCAGTGGATCTGGAACAGATTTCACTCTCACCATCAGCAGTGTGCAGGCTGAAGACCTGGCAGTTTATTACTGTCAGAATGATTATGTTTATCCGCTCACGTTCGGTGCTGGGACCAAGCTGGAAATAAAACGGTAATACTAGTAGCGGCCGCTGCAG
Anti-c-Myc (9E10)
GAATTCGCGGCCGCTTCTAGATGCAGGTGCAGCTaCAGGAGTCTGGGGGAGACTTAGTGAAGCCTGGAGGGTCCCTGAAACTCTCCTGTGCAGCCTCTGGATTCACTTTCAGTCACTATGGCATGTCTTGGGTTCGCCAGACTCCAGACAAGAGGCTGGAGTGGGTCGCAACCATTGGTAGTCGTGGTACTTACACCCACTATCCAGACAGTGTGAAGGGACGATTCACCATCTCCAGAGACAATGACAAGAACGCCCTGTACCTGCAAATGAACAGTCTGAAGTCTGAAGACACAGCCATGTATTACTGTGCAAGAAGAAGTGAATTTTATTACTACGGTAATACCTACTATTACTCTGCTATGGACTACTGGGGTCAAGGAGCCTCAGTCACCGTCTCCTCAGCCAAAACAACACCCAAGCTTGAAGAAGGTGAATTTTCAGAAGCACGCGTAGATATCGTTCTCACTCAATCTCCAGCTTTCTTGGCTGTATCTCTAGGACAGAGGGCCACCATCTCCTGtAGAGCCAGCGAAAGTGTTGATAATTATGGCTTTAGTTTTATGAACTGGTTCCAACAGAAACCAGGACAGCCACCCAAACTCCTCATCTATGCTATATCCAACCGAggttctGGGGTCCCTGCCAGGTTTAGTGGCAGTGGGTCTGGGACAGACTTCAGCCTCAACATCCATCCTGTAGAGGAGGATGATCCTGCAATGTATTTCTGTCAGCAAACTAAGGAGGTTCCGTGGACGTTCGGTGGAGGCACCAAGCTGGAAATCAAACGGGCTGTTGCTtaatactagtagcggccgctgcag
Anti-HuA
GTAAAACGACGGCCAGTgctagcCAAGTCCATTTAGTGGAGTCAGGTGGTGGCGTAGTTCAACCTGGAAGATCATTGACACTTTCCTGTGCTGCGTCCGGTTTTACCTTCAATTCTTATACGTTCCACTGGGTAAGACAAACCCCAGGTAAAGGTCTTGAATGGGTTGCAGTTTTAGCATACGACGGGAGTTACCAGCATTATGCAGATTCCGTGAAAGGAAGATTTACAATTTCCAGAGATAACAGCAAAAATACTTTATACTTGCAAATGAATAGTCTGAGACTTGAAGACACAGCGGTATACTTCTGTGCAAGGGGACAGACTACAGTTACTAAAATTGATGAGGACTATTGGGGGCAGGGTACATTGGTAATTGTCTCCTCCGGTGGTGGCGGTAGTGGAGGCGGTGGCTCCGGGGGAGGGGGCTCTGAGATCGTCCTTACCCAGTCTCCAGCAACGCTTAGTTTGTCTCCTGGAGAAAGAGCTACACTTTCTTGCAGAGCAAGCCAGTCTGTCTCTTCTTACTTAGCTTGGTATCAGCAAAAACCAGGTCAGGCTCCAAGGTTGCTGATATATGATGCATCTAACAGGGCGACTGGTATTCCTGCCAGGTTTTCCGGCTCCGGTTCTGGCACTGACTTCACACTTACTATTTCATCTCTAGAACCAGAAGATTTTGCCGTTTATTACTGCCAGCAGAGATCTAATTGGCCTAGGTCTTTTGGTCAGGGTACGAAGGTCGAGATCAAAAGGggatccGTCGTGACTGGGAAAAC
The sequences were cloned into the YD vector using BamHI and NheI restriction sites.
Construction of Expression Cassette
Sequence of the Aga1 protein was amplified from yeast genome using Q5 polymerase and sequences of the pTv3 promoter and CYC terminator was amplified from plasmids provided by Module 1 (working on synthetic haploids). The primers we used are disclosed in the table below.
| Name | Sequence |
|---|---|
| pTv3 down | TTAACAGAATTCGGATTCTCACAATCCTGTCGGT |
| pTv3 up | ATAATTCTCGAGAAGCTTGGACTTCCCACCGCCTTC |
| CYC TT down | ACAAAAGTCGACTCTAGACGAGCGTCCCAAAACCT |
| CYC TT up | ATATTAGAATTCATTATTAAATGGATCCTCATGTAATTAGTTATGTCACGCT |
| Aga1 down | ATATATGGATCCTTAACTGAAAATTACATTGCAAGCA |
| Aga1 up | ATATTAGAATTCACCACCATGACATTATCTTTCGCTCATTTTAC |
The amplification of Aga1 and assembly of the promoter and the terminator in one sequence was successful as proved by gel electrophoresis and subsequent gel purification. However, even after many repeated tries to clone the premature cassette into the pRS413 and pRS415 plasmids, our efforts remained futile. We tried using different ligases, amount of electroporated cells, and even a different cloning vector. As a result, we turned our focus solely on testing and improving the first of the three approaches - the original yeast display system.
Construction of Yeast Display Using only One Plasmid
This alternative greatly appealed to us thanks to its elegance and simplicity, but proved to be quite time-consuming to prepare. Therefore we only finished the design of the system that can be seen in Figure 3.4 below and assigned the idea only for future experiments.
Validation
Materials and Methods
Strains
E. coli
- DH5α
S. cerevisiae
- EBY100
Materials
- LB-M agar plates with ampicillin
- 1.5 ml eppendorf tubes
- 0.5 ml PCR tubes
- 50 ml centrifuge tubes
- NucleoSpin Plasmid DNA, RNA, and protein purification Kit
- NucleoSpin Gel and PCR Clean-up Kit
- LB-M medium with chloramphenicol
- YPD medium
- SD-min medium + CAA
- NaOH agarose gel and buffer
- TAE agarose gel and buffer
Methods
- Transformation
- Minipreparation of plasmid DNA
- Restriction digest
- Ligation
- NucleoSpin gel clean-up
- NucleoSpin plasmid DNA purification
- Streptavidin YD induction
- Yeast antibody labeling
Detailed description of all methods can be found on our Protocols page.
Results
Microscopy
Microfluidics
References
- ↑ Zhao, H.,Shen, Z.-M., Kahn, P. C. and Lipke, P. N. (2001). Interaction of α-Agglutinin and a-Agglutinin, Saccharomyces cerevisiae Sexual Cell Adhesion Molecules. Journal of Bacteriology, 183(9), 2874–2880. doi:10.1128/JB.183.9.2874-2880.2001
- ↑ Lipke, P. N. and Kurjan, J. (1992). Sexual agglutination in budding yeasts: structure, function, and regulation of adhesion glycoproteins. Microbiology and Molecular Biology Reviews, 56(1), 180-194.
- ↑ Boder, E. T. and Wittrup, K. D. (1997). Yeast surface display for screening combinatorial polypeptide libraries. Nature Biotechnology, 15, 553-557. doi:10.1038/nbt0697-553
- ↑ Baird, C. et al. (2012). Yeast Display scFv Antibody Library User’s Manual. Richland, WA: Pacific Northwest National Laboratory.
- ↑ Wasilenko, J. L. et al. (2010). Cell Surface Display of Highly Pathogenic Avian Influenza Virus Hemagglutinin on the Surface of Pichia Pastoris Cells Using alpha-Agglutinin for Production of Oral Vaccines. Biotechnology Progress, 26(2), 542-547. doi:10.1002/btpr.343.

