The main goal of our project is to prevent accidental dissemination in the environment of GEO, and more specifically, of Escherichia coli, the most widely used chassis in iGEM. But before elaborating sophisticated physical and biological containment systems to prevent dissemination, we thought it would be interesting to study the E. coli survival in different natural environments.
Medium
To count our E. coli strains, we chose to use the Mac-Conkey medium. This medium is very selective for Gram negative enteric bacilli, so Escherichia coli. It contains bile salt and crystal violet dye to inhibit Gram positive strains, and neutral red dye which turns pink all colonies able to fermentate the carbon source (for the first test it was lactose, and after sorbitol).
Thus, by adding the appropriate antibiotic to this medium, we thought we would be able to select our strains only, although we needed to verify this hypothesis.
Preliminary Study
Our experiment relies on the hypothesis that we will be able to detect selectively our E. coli strain when plating a sample taken from our tested environments on MacConkey plates supplemented with the appropriate antibiotic. We therefore needed to test this hypothesis and verify that the natural microorganisms found in our environment samples would grow or not on our selective plates. We also decided to grow our samples on LB medium as this medium is very rich and allows the growth of many microorganisms. We wanted to see if there was a difference with the MacConkey (MCK) medium, and if we could see the growth of the natural "inhabitants" of our samples.
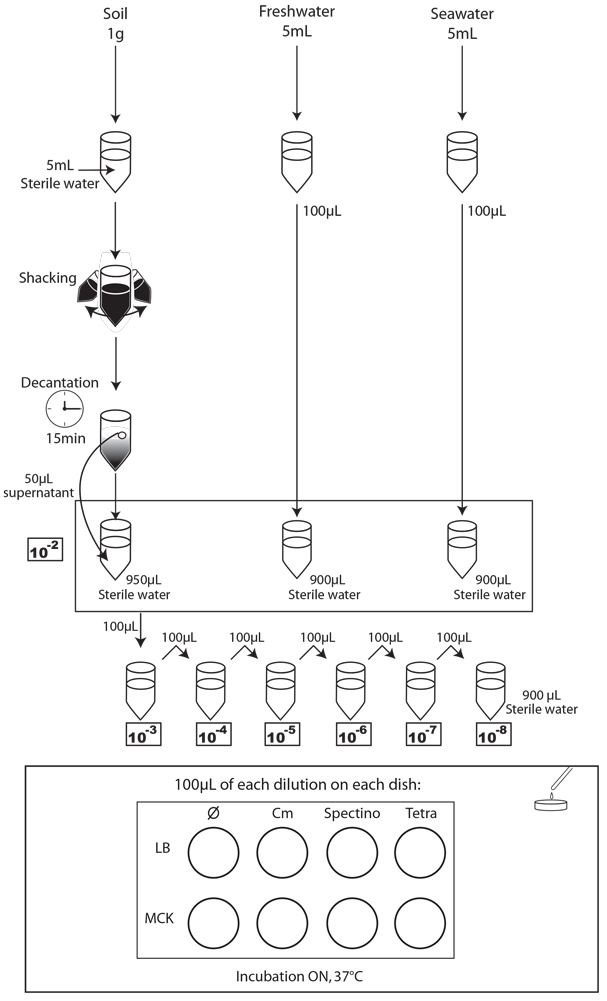
We chose to analyze the soil, seawater and freshwater samples. We prepared 5 dilutions (10-2 to 10-6) of a sample of our natural environment and plated 100 µl on the following media:
- LB
- LB + Spectinomycin (Spectino : 100ng/μL)
- LB + Tetracycline (Tetra : 10ng/μL)
- LB + Chloramphenicol (Cm : 20ng/μL)
- MCK
- MCK + Spectinomycin (Spectino : 100ng/μL)
- MCK + Tetracyclin (Tetra : 10ng/μL)
- MCK + Chloramphenicol (Cm : 20ng/μL)
We next incubated the plates overnight at 37°C and observed them the following day.
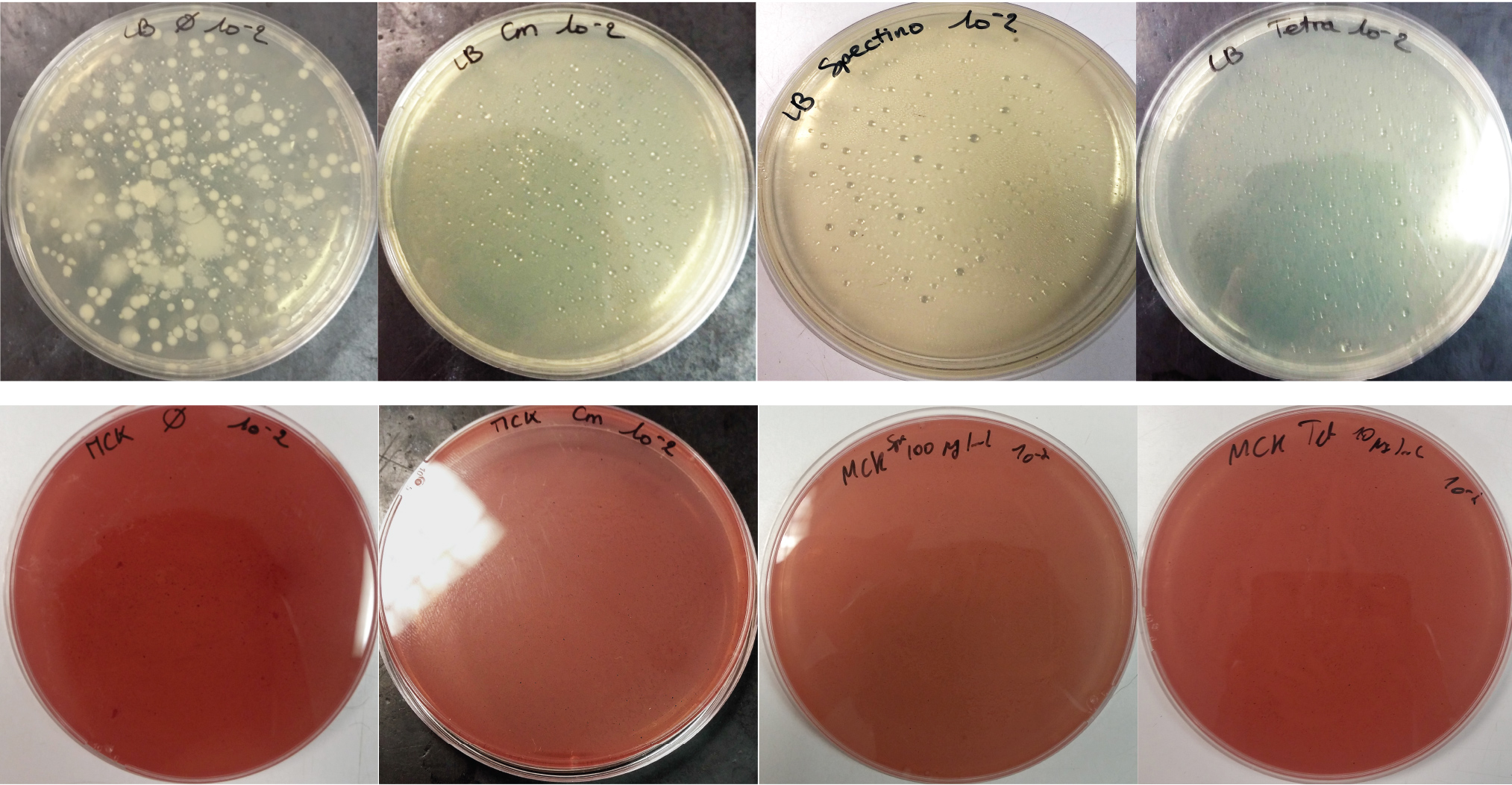
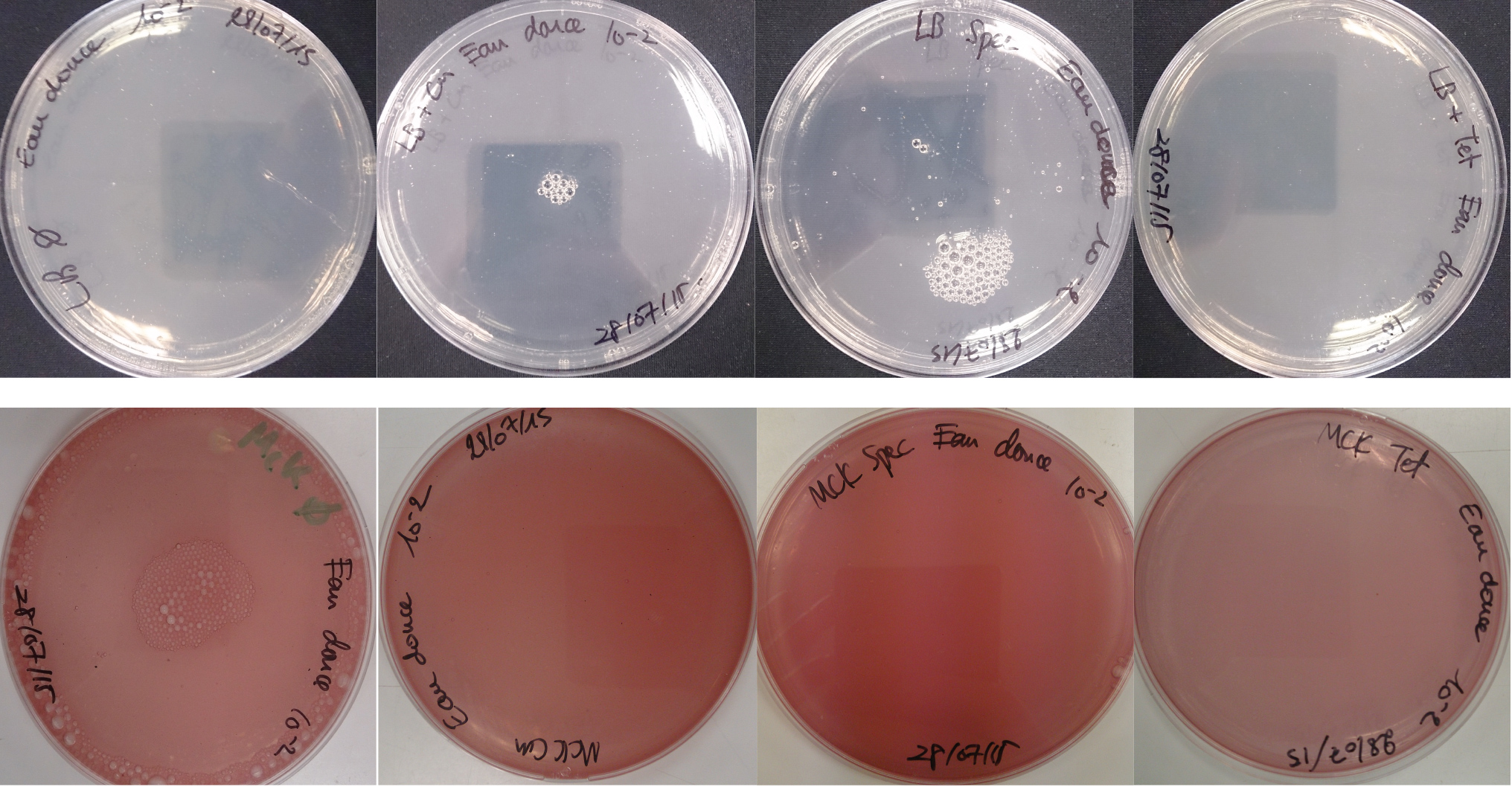
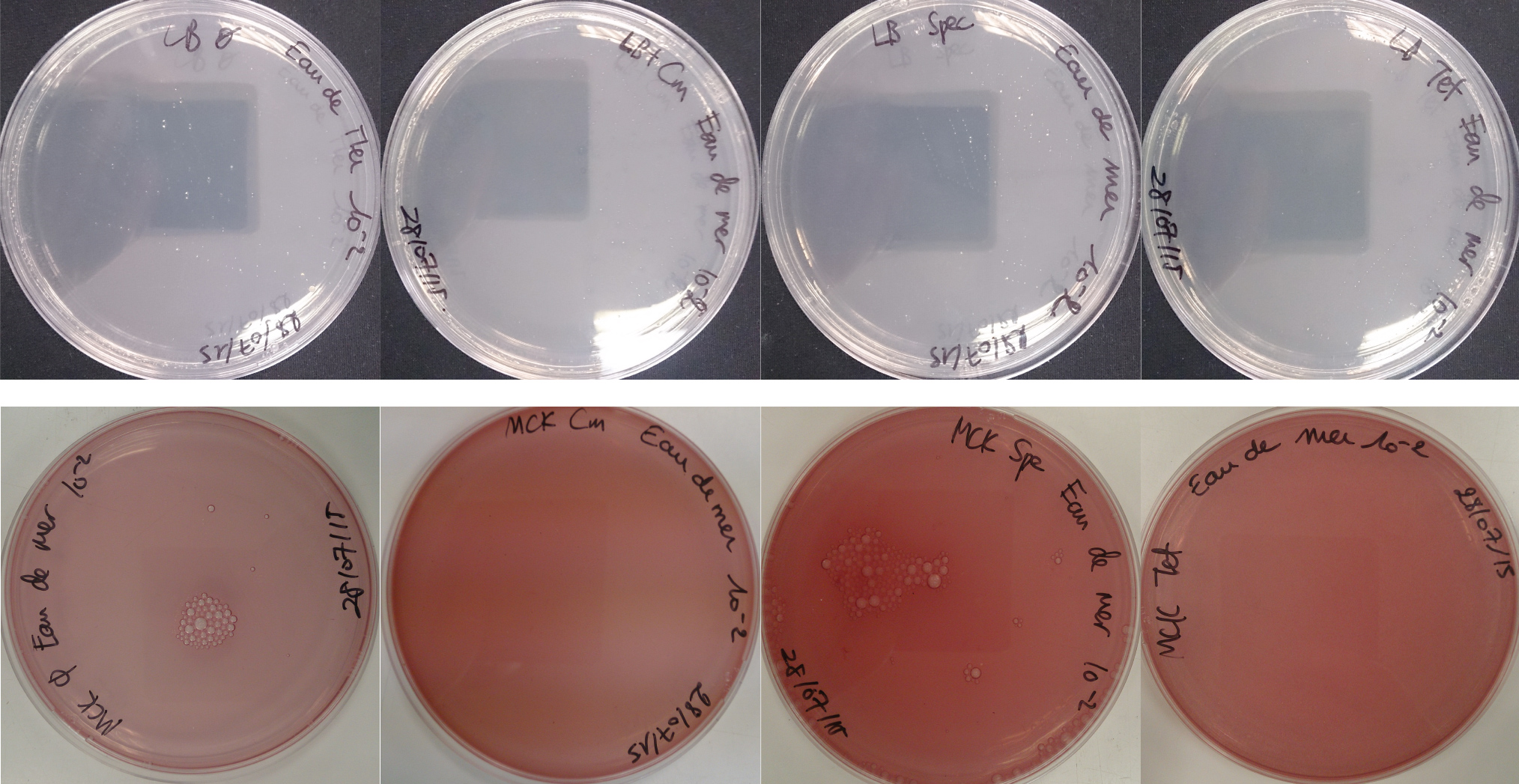
For the soil experiment Figure 1.1, we observed the growth of microorganisms on LB medium for the 10-2 dilution. These micro-organisms, however, appear to be sensitive to three antibiotic tested (chloramphenicol, spectinomycin and tetracycline). They are also unable to grow on the MacConkey medium. No growth of micro-organisms was observed when samples of freshwater Figure 1.2 or sea water Figure 1.3 were plated on either LB or MacConkey media. This may have been expected for the seawater medium, as micro-organisms from this environment may require high salt media for growth.
Chiffres populations bactériennes dans les différents environnement
On the whole, we did not observe growth of “natural” microorganisms on the MacConkey + antibiotics plates, demonstrating that in the conditions tested, the only micro-organisms that we should observed are our specific lab bacteria strains.
Figure 1.1 - Preliminary study for soil environment - Test of soil natural microorganisms growth on LB or MCK with or without Chloramphenicol, Spectinomycin and Tetracycline. Soil sample diluted 10-2
Figure 1.2 - Preliminary study for freshwater environment - Test of freshwater natural micro-organisms growth on LB or MCK with or without Chloramphenicol, Spectinomycin and Tetracycline. Freshwater sample diluted 10-1
Figure 1.3 - Preliminary study for seawater environment - Test of seawater natural micro-organisms growth on LB or MCK with or without Chloramphenicol, Spectinomycin and Tetracycline. Seawater sample diluted 10-1
Study
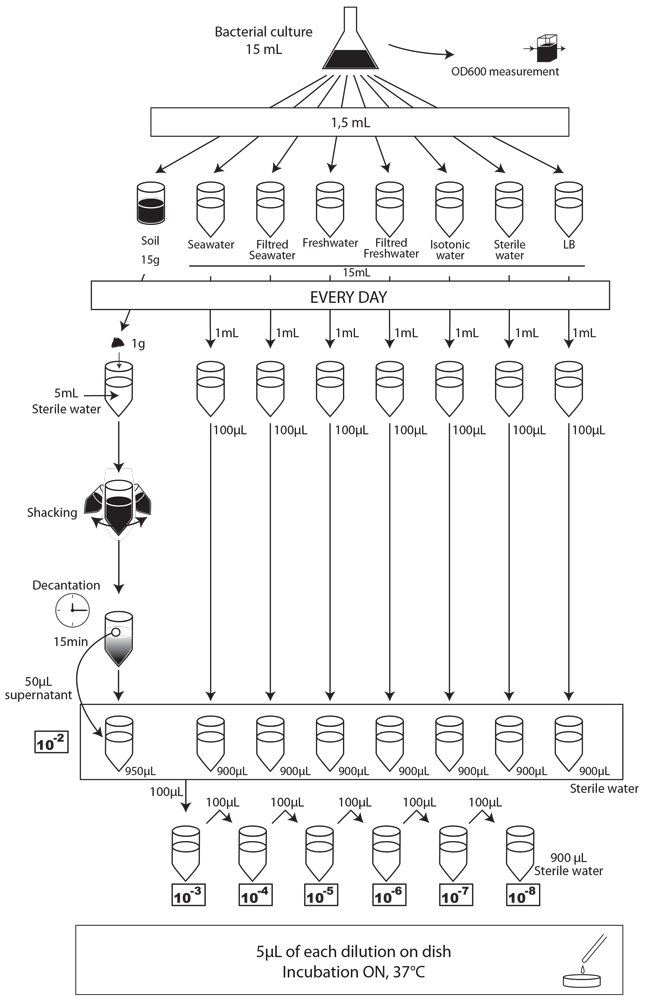
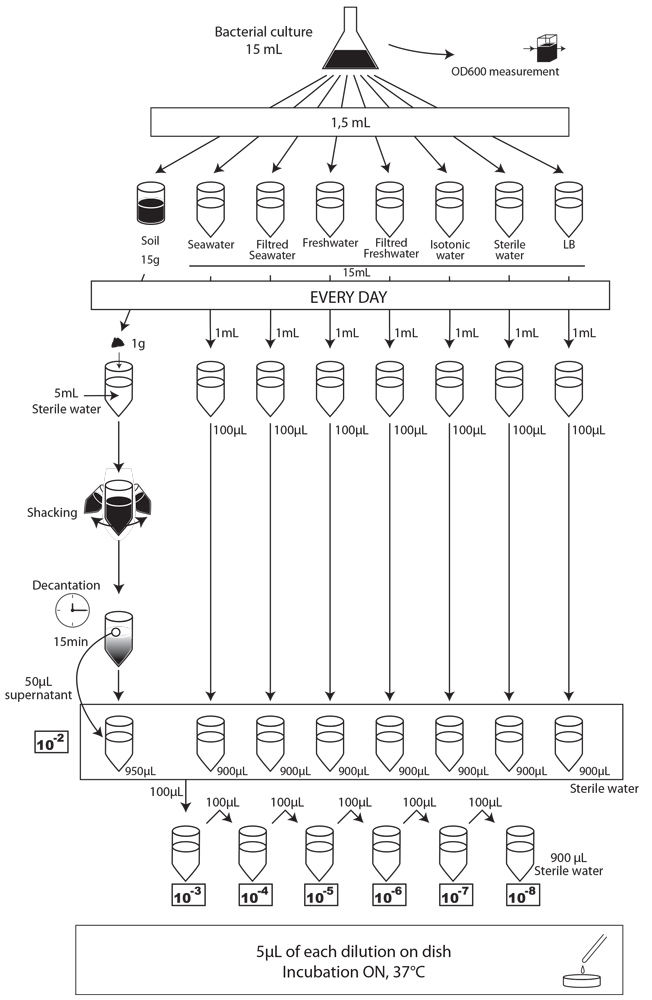
To simulate an environmental contamination by GEOs, we introduced a bacterial culture grown in LB without any antibiotic in our environment samples. Indeed, in case of an accidental dissemination, what could happen to our strain if we spill a bacterial culture into the wild? We chose to use the same protocol for each environment, except an additional step of water dilution for the soil sample (Figure XX):
- We prepared a 15mL culture of our three different strains (E. coli 1320, 1693 and 1696), in LB medium, without antibiotic the day before starting the experiment.
- The first day (D0), we measured the optical density at 600 nm (OD600) to evaluate “concentration” in bacteria , and we plated on MCK with the appropriate antibiotic a sample of each culture to count the CFU.
- We put then 0,5mL of each of the three cultures in each of the eight experimental environment (5mL for liquid environment, 5g for soil), and shake it to make a homogeneous mix. Negative controls (-) are environmental samples without culture contamination. All experimental samples were left on the bench, at room temperature, in a 15mL closed falcon tube.
- Each day, we took a 1mL sample of each contaminated liquid experimental environment and prepared dilutions of this sample from 10-1 to 10-8 in isotonic water. For the soil environment, we took 1g of soil, which we diluted in 5mL of sterile water, and after one step of shaking and of 15 min decantation, we pipetted 50 µL of the supernatant and diluted it in 950 µL of isotonic water. This allowed us to obtain the 10-2 dilution. We prepared then the 10-3 to 10-8 dilutions as described for the liquid environments.
- We spotted 5 µL of each dilution on the corresponding MCK plate + antibiotic (no antibiotic for the control).
- We incubated of plates overnight at 37°C.
- The next day, we counted the CFU to estimate our strain survival.
Results
It took us time to develop a proper protocol to follow the survival of our strains in the various environments. It was a trial and error process and we had to adjust the protocol after each trial.
Preliminary - experiment
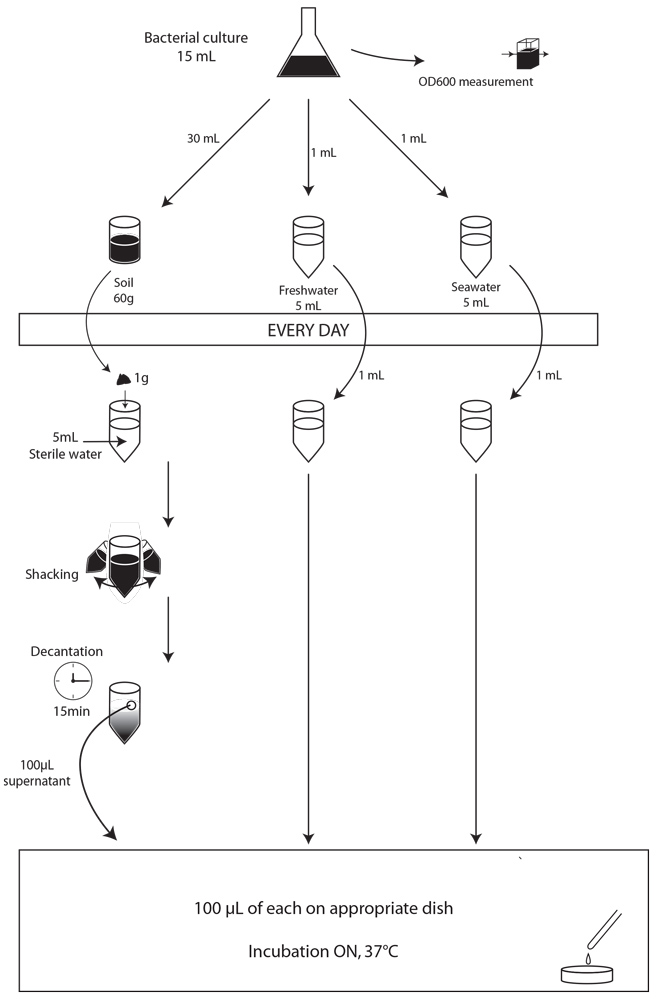
First of all, we made a small-scale preliminary experiment during three days to determine whether our strains were able to survive in our three major environments: soil, freshwater and saltwater, and to determine what would be the optimal concentration of bacteria in our contaminating culture for our experiment. We used 5g of soil, and 5mL of freshwater or salt water. We contaminated them with 1mL of overnight bacterial culture not diluted or diluted 10 or 100 times. For these preliminary experiments, the samples to be plated were prepared as described above, but rather than spotting 5 µl on plates, we plated 100 µl of the dilutions on the adequate MCK plates and incubated the plates at 37°C overnight. Unfortunately, because of some problems with the medium, the results for the experiments with freshwater and saltwater could not be interpreted. The results for the soil experiment are presented Figures X, Y , Z
|
D0 |
D1 |
D2 |
D3 |
| CM 1 |
more than 1000 |
186 |
111 |
141 |
| CM0,1 |
1744 |
35 |
13 |
2 |
| CM 0,01 |
272 |
1 |
0 |
0 |
| Spe 1 |
more than 1000 |
490 |
424 |
276 |
| Spe 0,1 |
more than 1000 |
67 |
75 |
88 |
| Spe 0,01 |
441 |
0 |
1 |
2 |
| Tet 1 |
more than 1000 |
1308 |
524 |
416 |
| Tet 0,1 |
more than 1000 |
196 |
35 |
62 |
| Tet 0,01 |
414 |
0 |
0 |
0 |
| control |
0 |
1 |
3 |
4 |
On the whole, we observed that after three days of contaminations only, there were no or very few surviving E. coli bacteria in the soil when we diluted our contaminating E. coli culture 10 or 100 times. Moreover, the same decrease trend is observed for each of the three E. coli strains tested, suggesting that the integration of each of the three resistance genes in the genome of the E. coli MG1655 strain is not introducing a bias in our experiment. From this experiment, we concluded that it was more interesting for us to use an undiluted bacterial culture to contaminate our environment in our following experiments.
First Experiment
We used the same protocol than before, but with 30g of soil and 30mL of water. We made this experiment in triplicate.
Soil
We started a new experiment with more soil (30g) to analyze the contamination evolution during a longer period of time. We planned to do it for a few weeks but after three days, we observed contaminations on our negative control. After multiple trials, we concluded that this was due to the MCK medium used. Thus we changed the MCK stock and did tests from which we concluded that adding sorbitol to the MCK was more selective than adding lactose. Consequently, we decided to start another time the experiment for all the different contaminated media using Sorbitol-MCK rather than lactose-MCK.
Seawater and Freshwater
The first time we put our strains in 5mL of water but we encountered two main problems. The first one was that it was very difficult to spread the 100µL sample on plates. The samples seemed to slide on the medium. And the next day, we can only observe some “lines” of colonies. The second problem appeared on the second day: the medium, normally red, turned into a yellow color. This was probably due to a change of pH in the water sample, most likely due to the metabolism of some microorganisms when adding LB in the environment.
Final experiment
After those pre-tests, we finally had established a protocol that should allow us to carry out our experiment. We chose to test 15g (soil) or 15mL (seawater and freshwater) of our samples contaminated by 1,5mL of our bacterial cultures. We used the following protocol.
We performed CFU analyses each day by depositing 5µL spots of contaminated media on specific MCK plates. The results are presented Figure XXX (A to I).
Due to the lack of time, we were not able to conduct these experiments in triplicates and for a longer period of time, which would be required to obtain solid results from which conclusions can be drawn. Nonetheless, these experiments allowed us formulate to some preliminary conclusions. First, in our negative control (corresponding to samples of uncontaminated environments plated on MCK plates without antibiotics), we observed a very limited number of bacteria in all our samples except for the freshwater sample. This number also appeared to be stable in time, suggesting that the population of the micro-organisms that grew on our plates remained stable during the time of our experiment. [explication pour freshwater?].
Before interpreting the results of our contamination experiments, it is important to remember that when we did not observed the growth of any micro-organisms from the various environments tested when antibiotics were added to the MacConkey medium.
Physical containment
Considering the non-negligible risk of escape of a GEO, we wanted to add an extra layer of safety in our project. Therefore, we devised a way to physically contain E. coli and other bacteria without hindering it to perform its main function. Indeed, the system we designed is a porous glass beam in which bacteria can grow, survive and carry out their function. The nutrients of the external medium, like sugars, amino acids, etc. can penetrate inside the devise but the bacteria cannot escape from it.
Thus, the SafetE.coli chassis can be in a safer manner in contact with natural environments, which is an important aspect when the project competes in iGEM competition tracks such as Environment or Health and Medicine.
Some teams include in their project a devise, more or less sophisticated, in which the GEO or their products will perform its function. Here we can cite the 2014 project of NCTU Formosa where we can found an example of such devise: The Pyramid Trap containing PBAN (Pheromone Biosynthesis Activating Neuropeptide) produced by a Genetically Engineered Escherichia coli. However, other teams do not have the time or means to do as well.
Thus, the 2015 Paris Saclay team project offers to all iGEM teams a fast, efficient and inexpensive way to build an impenetrable shelter for bacteria.1 Mettre en note de bas de page: (1) Mercedes Perullini, Mat´ıasJobba´gy, Galo J. A. A. Soler-Illia, and Sara A. Bilmes, Chem. Mater. 2005, 17, 3806-3808
The system principle
The aim of the containment is to create a physical barrier which should prevent accidental spreading of bacterial culture. The idea is to make a containment where bacteria can grow but which does not allow them to be in contact with the environment. Bacteria will be contained inside silica monoliths. To enable cell growth, we found a protocol which describes how to make cavities inside silica monoliths possible.
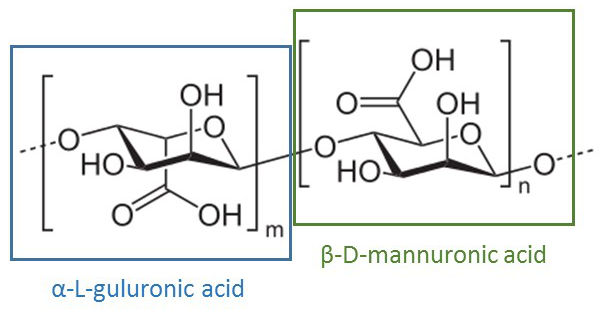
First, cells are encapsulated in an alginate gel. The alginic acid polysaccharide is a linear copolymer of β-D-mannuronic and α-L-guluronic acid extracted from brown algae or bacteria.
Alginic acid polysaccharide
The bacterial culture is mixed with sodium alginate and the solution is dropped in a chloride calcium solution. At neutral pH, carboxylic acid functions are deprotonated so that the polymer bears a global negative charge, usually compensated by sodium ions. Addition of divalent cations such as Ca2+ induces cross-linking of the polymer, and therefore gel formation.
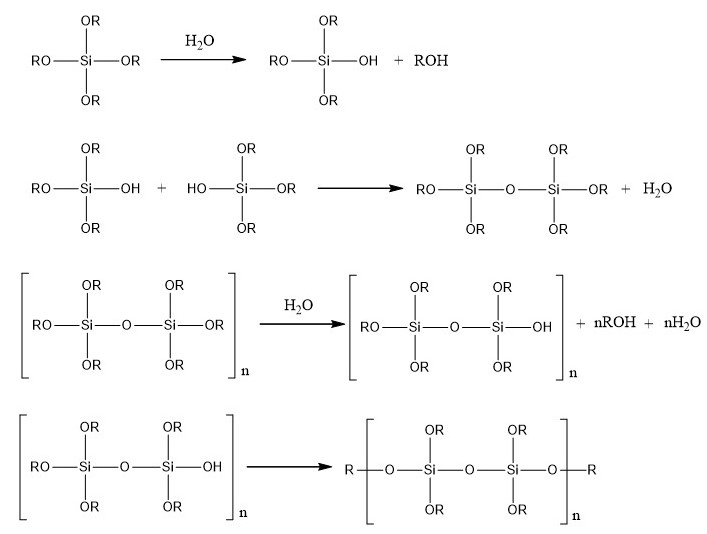
Then, silica monoliths are created around the beads obtained by a sol-gel process which allows the formation of silica gels at room temperature from aqueous precursors (Brinker and Scherrer 1990). The reactions involved in this process are presented in the figure below.
FPolymerization reactions involved in the sol-gel process
Finally, the gel is dissolved by using an acid which chelates Ca2+ so that cells have a cavity where they can grow and which can be filled with a defined growth medium.
Protocol
The first one is performed by dropwise addition of a 1.5% (w/w) sodium alginate cells suspension in a 0.1 M CaCl2 solution. After 10 min stirring, about 3 mm diameter beads are easily collected by filtration. The calcium alginate polymer prevents cell contact with synthesis precursors. The second step consists of silicate polymerization in the presence of commercial silica nanoparticles (Ludox HS40 from Aldrich), leading to a nanoporous monolithic structure. Monoliths are prepared at room temperature by mixing 2 volumes of 1.25 M sodium silicate with 1 volume of colloidal silica and 1 volume of succinic acid (5 wt %) into a recipient containing the alginate-cells bead. Once the sol-gel polymerization reaction is completed, the stiff monolith obtained is left in contact with 0.05% potassium citrate 3h. To provide necessary nutrients to the immobilized cells, potassium citrate solution is further replaced by LB medium according to encapsulated cell strain requirements.1,2 Mettre en note de bas de page: (2) Mercedes Perullini, Frédéric Orias, Claude Durrieu, Matias Jobbagy, Sara A. Bilmes, Biotechnology Reports, 2014, 4, 147-150
Results
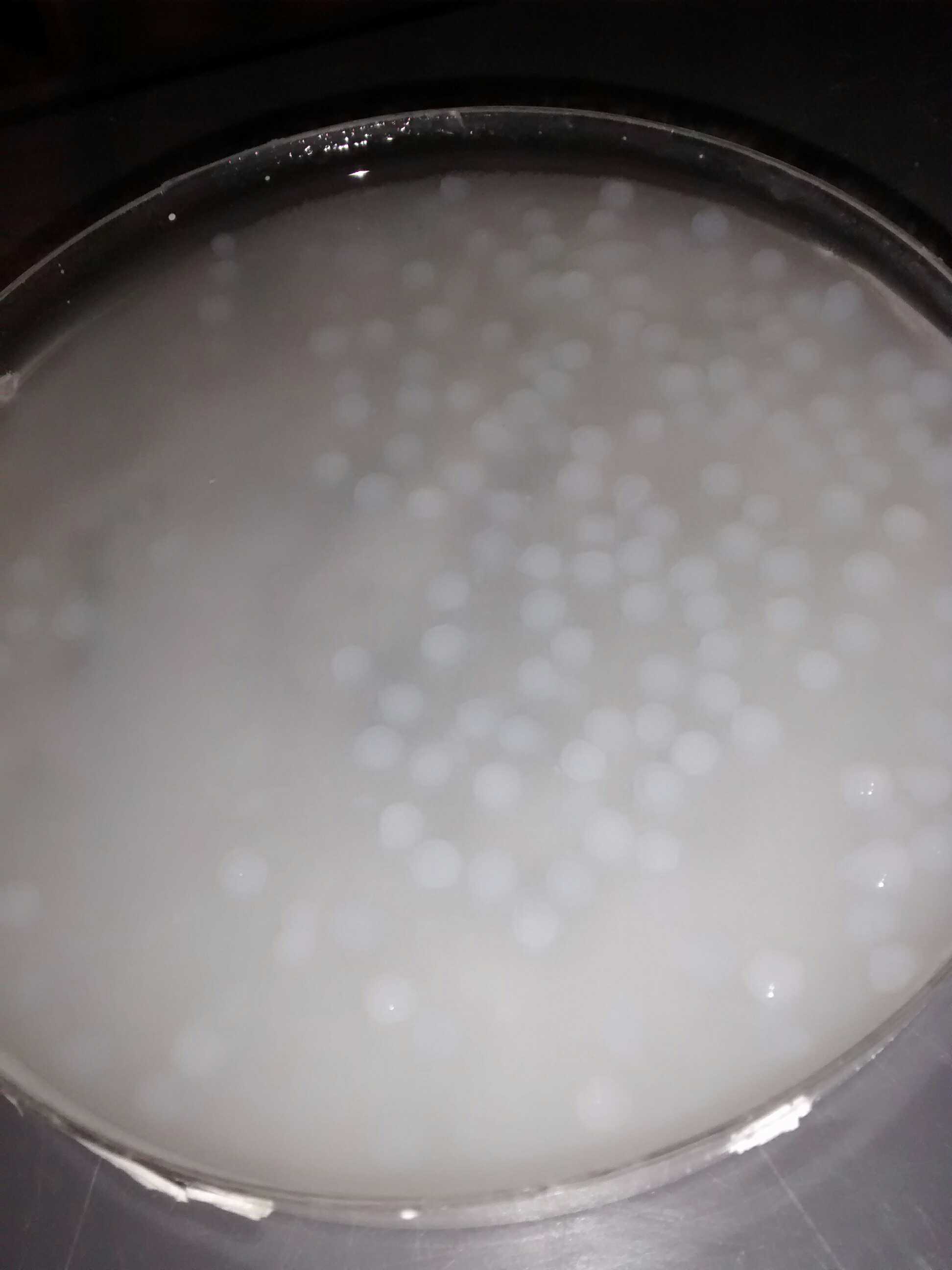
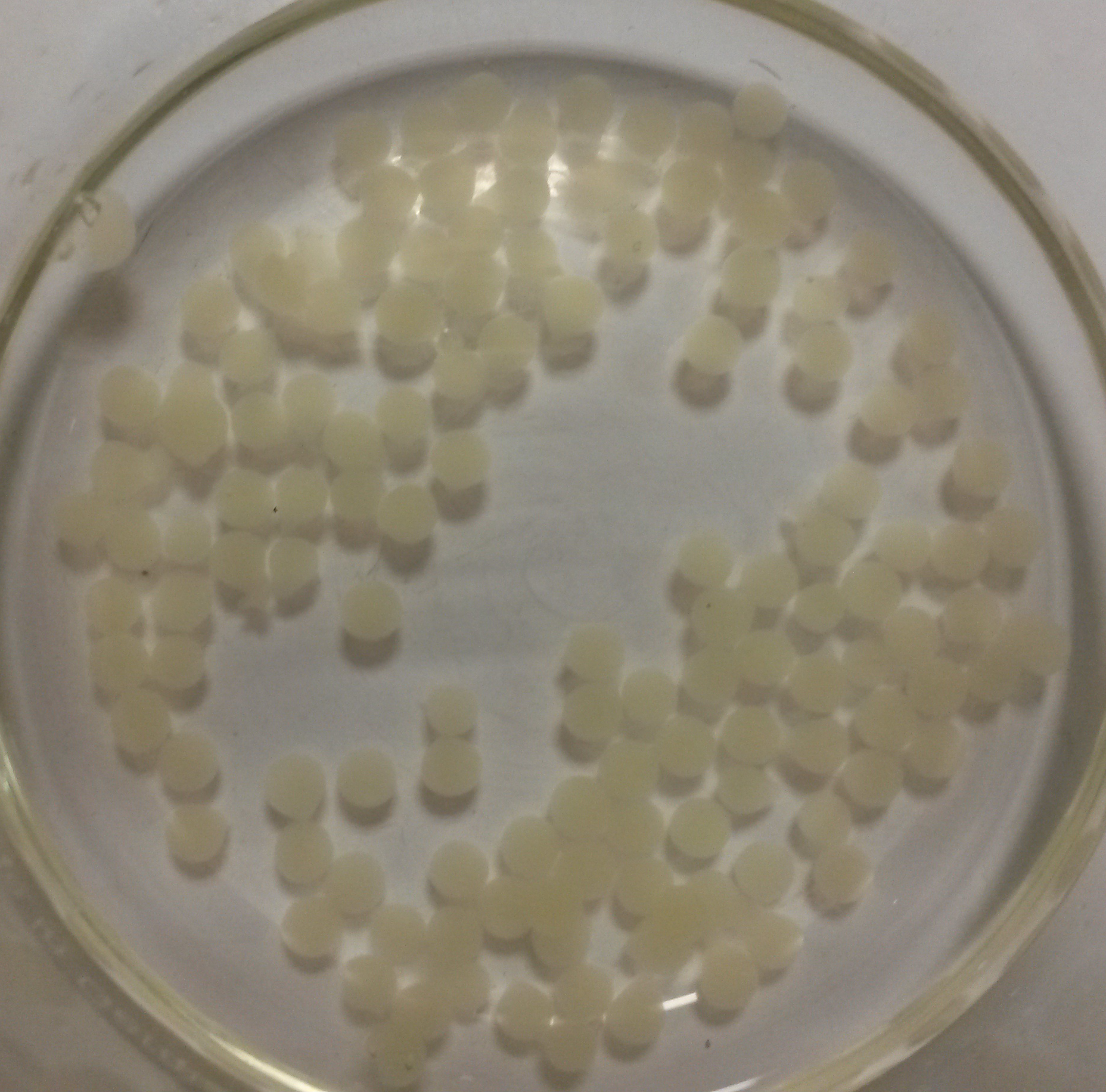
We obtained nice beads which are properly jellified and measure about 3 mm as expected.
Alginate beads of bacterial culture
The silica monolith was obtained only when we let it in contact with air.
Silica monolith with cavities containing bacterial culture
We did not have the time to test our system. It will be interesting to see if bacteria do not escape from the silica monolith and if they survive inside. The Aachen team suggested us different devices to measure the growth rate. The first one can be used to determine biomass concentration using capacity measurements.3 The second one measures growth rate using an optical detection system that is based on measuring the intensity of back-scattered light from bacterial cells suspended in the liquid culture.4 Mettre en note de bas de page: (3) Noll, T., & Biselli, M., Journal of biotechnology, 1998, 63, 187-198. (4) Toprak, E., Veres, A., Yildiz, S., Pedraza, J. M., Chait, R., Paulsson, J., & Kishony, R., Nature protocols, 2013, 8, 555-567.
The Temperature-Based System for E. coli: SafetE.coli
Perspectives
Modeling