Team:Waterloo/Modeling/CaMV Biology
Cauliflower Mosaic Virus (CaMV) Biology
Discuss briefly why we chose CaMV (double-stranded) and common, similar to cassava mosaic virus and other important guys
Stuff from Viral Replication Page
Overview of Genome
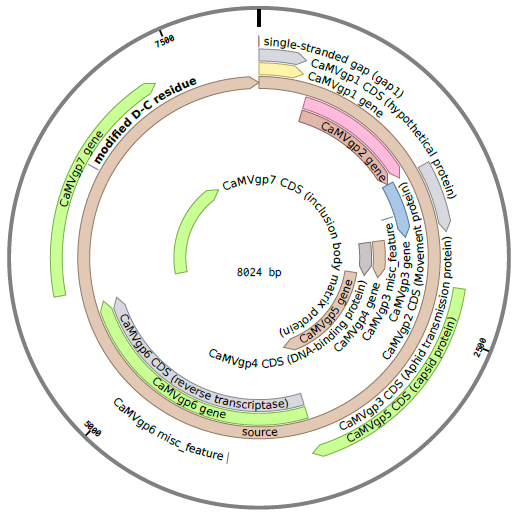
The CaMV genome consists of circular, double-stranded DNA approximately 8000 bp in length . The genome contains three discontinuities typical of pararetroviruses . Discontinuities are repaired within the nucleus to form covalently closed circular (ccc) DNA . There are seven to eight open reading frames (ORFs) which code for six proteins of known function (P1-P6) . Two main RNAs are produced from cccDNA: pregenomic 35S RNA and subgenomic 19S RNA . A third type of transcript, 8S RNA, is also described below, but not incorporated in our model.
Pregenomic 35S RNA
35S RNA is a complete transcript of the CaMV genome with additional 180 nt terminal repeats . It has two primary functions. The first is to act as an intermediate in genome replication. That is, it's reverse transcribed either in a viral factory or within the capsid to recover gapped dsDNA . It's second function is as polycistronic mRNA coding for P1-P5. In order to produce P1-P5 the 35S promoter must be transactivated by P6 . 35S RNA also undergoes alternative splicing to regulate P1 and P2, which are potentially toxic in high enough concentrations . As such, we account for the relative concentrations of spliced and unspliced 35S RNA in our model since only unspliced 35S RNA will be encapsidated.
Subgenomic 19S RNA
The 19S RNA is a 2.3 kb transcript which contains the ORF for P6 . P6 is constitutively produced from 19S RNA and it's many functions are described in it's own section. Other than producing P6 it has no other function.
8S RNA
8S RNA is 610-680 nt long and is transcribed from the leader sequence of 35S RNA. It's believed that it serves as a decoy for RNA interference, which normally targets the 35S leader . Since it's non-coding and is not targeted by our CRISPR system we do not track 8S RNA in our model.
Movement Protein (P1)
The movement protein (MP) of CaMV facilitates intercellular spread of virions . P1 binds to pectin methyltranferase and ultimately forms intercellular tubules though which virions are transported to neighbouring cells . This protein may also bind cooperatively to RNA to direct 35S and/or 19S RNA to neighbouring cells , which suggests the protein-RNA complex may also move between cells without a capsid. As we are not modelling in detail the transport of virions outside the cell we do not track P1 concentration in our model. Rather, the rate of virions exiting the cell is assumed to be proportional to the number of virions. Additionally, we do not model the possible transport of 35S or 19S RNA transported outside the cell through these tubules.
Aphid Transmission Factor (P2)
The aphid transmission factor (ATF) is a protein which allows for host-to-host transmission of the virus using aphids as vectors. P2 binds to both the aphid stylets and to P3 which allows virions to associate with the stylet upon feeding . It's primary function is host-to-host transmission so we do not track P2 concentration in the cell, but it does produce one electron-lucent inclusion body (elIB) which has minor effects on the system. To form the elIB, P2 associates with microtubules and are transported along the cytoskeleton to a central location. In this process some P3 associates with the P2 and is transported to the elIB . Finally, a small percent (~5%) of virions find their way to the elIB . In this model the amount of P3 and virions which find their way into the elIB are assumed to be negligible.
Virion-Associated Protein (P3)
The virion associated protein (VAP) is a 15 kDa protein is weakly associated with the viral capsid . It's implicated in both host-to-host and cell-to-cell transmission. Due to it's importance in transmission, we track P3 concentration over time and consider a virion "complete" only if it has P3 bound to it.
Capsid Protein (P4)
The capsid protein (CP) is a precursor to numerous other polypeptides which we refer to as P4 subunits. The P4 subunits include P37, P44, P40, P80, P90, and P96, with P80 and P96 being homodimers composed of P37 and P44 respectively . These subunits come together to form a capsid consisting of three layers and using 420 subunits . This model does not capture the complexity of P4 splicing and capsid assembly, rather it assumes packaging is proportional to the amount of P4 available in the cell.
Reverse Transcriptase (P5)
The reverse transcriptase (RT) is a protein which reverse transcribes 35S RNA to form partially double stranded (pds) DNA . The pdsDNA is then encapsidated by P4 subunits to produce virions. It's unknown if reverse transcription occurs within the capsid or in the viroplasm . There is evidence that in Hepatitis B (a similar pararetrovirus) that RT first binds to the pregenome, the RT-RNA complex is recognized by CP to form a capsid, then reverse transcription occurs within the capsid . In this model we assume the reverse transcription and packaging occur simultaneously.
Transactivator/Viroplasmin Protein (P6)
The transactivator/viroplasmin protein (TAV), as it's name suggests, is a protein primarily involved in transactivating the five other proteins and condenses into viroplasms . The viroplasms, aka viral factories or electron-dense inclusion bodies (edIBs), sequester 35S RNA transcripts and produces the other proteins required for viral spread and assembly . The purpose is two-fold: first of all it concentrates the nucleic acid and protein in a smaller volume to increase reaction rates, and secondly it protects the replication machinery from cell defenses . We do not model the formation of edIBs in this model, instead we assume all P6 accumulates as it's produced. This allows us to model the activation of P1-P5.
Here we present a brief overview of the CaMV replication cycle, compiled from Khelifa et al. 2010 (), Yasaka et al. 2014 (), Hohn, Fütterera & Hull 1997 (), and Citovsky, Knorr, & Zambryski 1991 ()
- The virion enters the host cell, moves to the nuclear pore, and the viral genome is imported in the nucleus
- Discontinuities in the genomes are repaired to form minichromosomes
- Minichromosomes are used as templates to produce 35S RNA, 19S RNA, and 8S RNA
- P6 is translated from 19S RNA and accumulates in electron-dense inclusion bodies (edIBs)
- P6 transactivates production of P1-P5 from 35S RNA
- Packaging occurs. The exact process is complicated, but either :
- 35S RNA is packaged into virions along with P5 then is reverse transcribed within the capsid, or
- 35S RNA is reverse transcribed in the edIBs before packaging
- The virion may either reinfect the nucleus to increase the copy number of the minichromosome (early in the infection) or it may leave the cell
Due to the complicated nature of viral packaging, we decided in our model to combine the packaging reaction into one term, dependent on P4, P5, and unspliced 35S RNA concentration.
Someone please update, maybe Laura?
Lifecycle Overview
Infection
Replication
Intercellular Spread
CaMV Genome Structure
Natural Plant Defenses
e.g. RNAi