Difference between revisions of "Team:Goettingen/Results"
| Line 312: | Line 312: | ||
<p> | <p> | ||
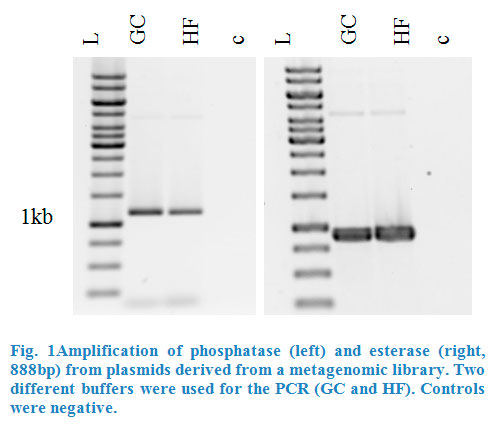
Esterase was amplified from pET101_E064 and phosphatase from pTOPOXL_PLP07 by PCR for each enzyme (Fig.1). Both plasmids were given to us by the Applied | Esterase was amplified from pET101_E064 and phosphatase from pTOPOXL_PLP07 by PCR for each enzyme (Fig.1). Both plasmids were given to us by the Applied | ||
| − | and Genomic Microbiology department. Primers contained restriction sites <em>Kpn</em>I and <em>Sac</em>I for esterase and | + | and Genomic Microbiology department. Primers contained restriction sites <em>Kpn</em>I and <em>Sac</em>I for esterase and <em>Xho</em>I and <em>Pst</em>I for the phosphatase in order to make |
them compatible for insertion into the multiple cloning site of the pBAD vector. The genes for these two enzymes had been found in screenings of | them compatible for insertion into the multiple cloning site of the pBAD vector. The genes for these two enzymes had been found in screenings of | ||
metagenomic libraries. | metagenomic libraries. | ||
| Line 347: | Line 347: | ||
</p> | </p> | ||
<p> | <p> | ||
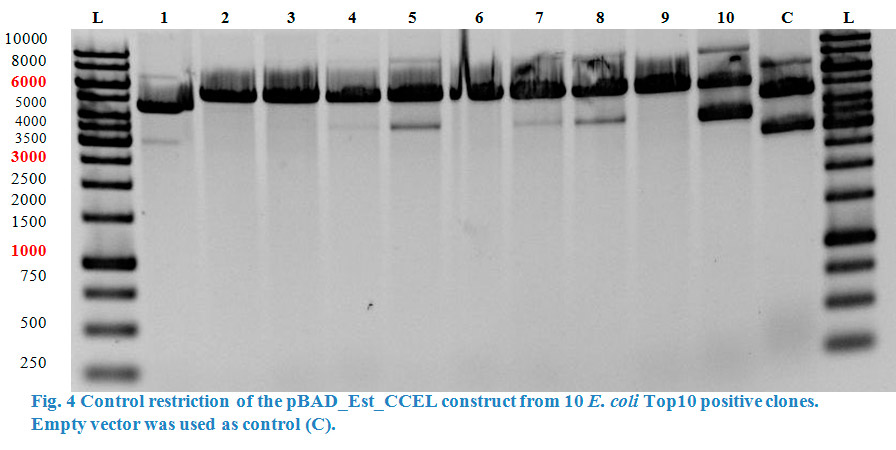
| − | A restriction control with | + | A restriction control with <em>Sac</em>I was run in a 0,8% agarose gel (Fig. 4). Samples #2, #3 and #9 correspond<strong> </strong>to the linear plasmid containing |
the insert. | the insert. | ||
</p> | </p> | ||
Revision as of 12:40, 18 September 2015
Project Results
Transformation Efficiency Kit, RFP construct (iGEM)
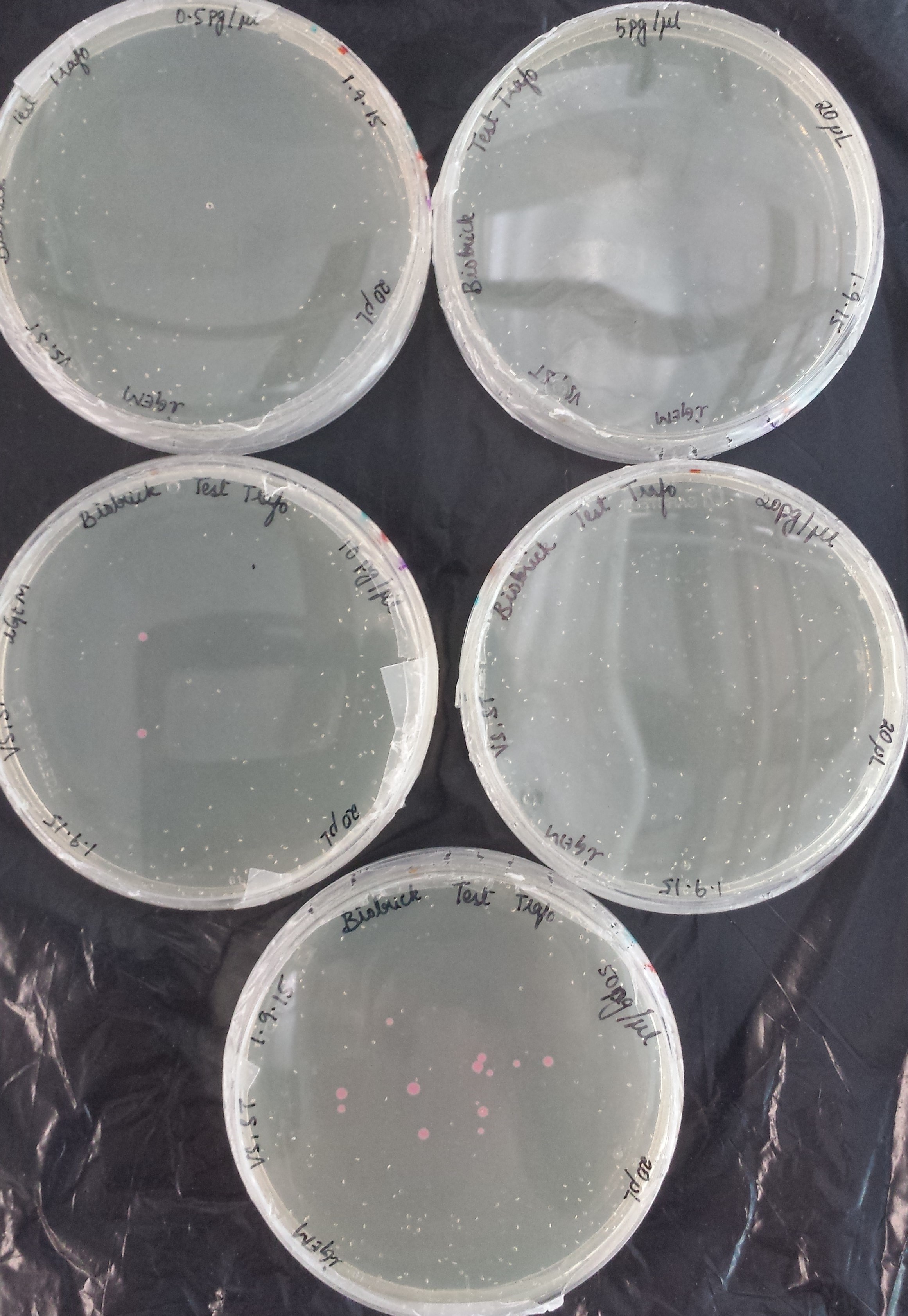
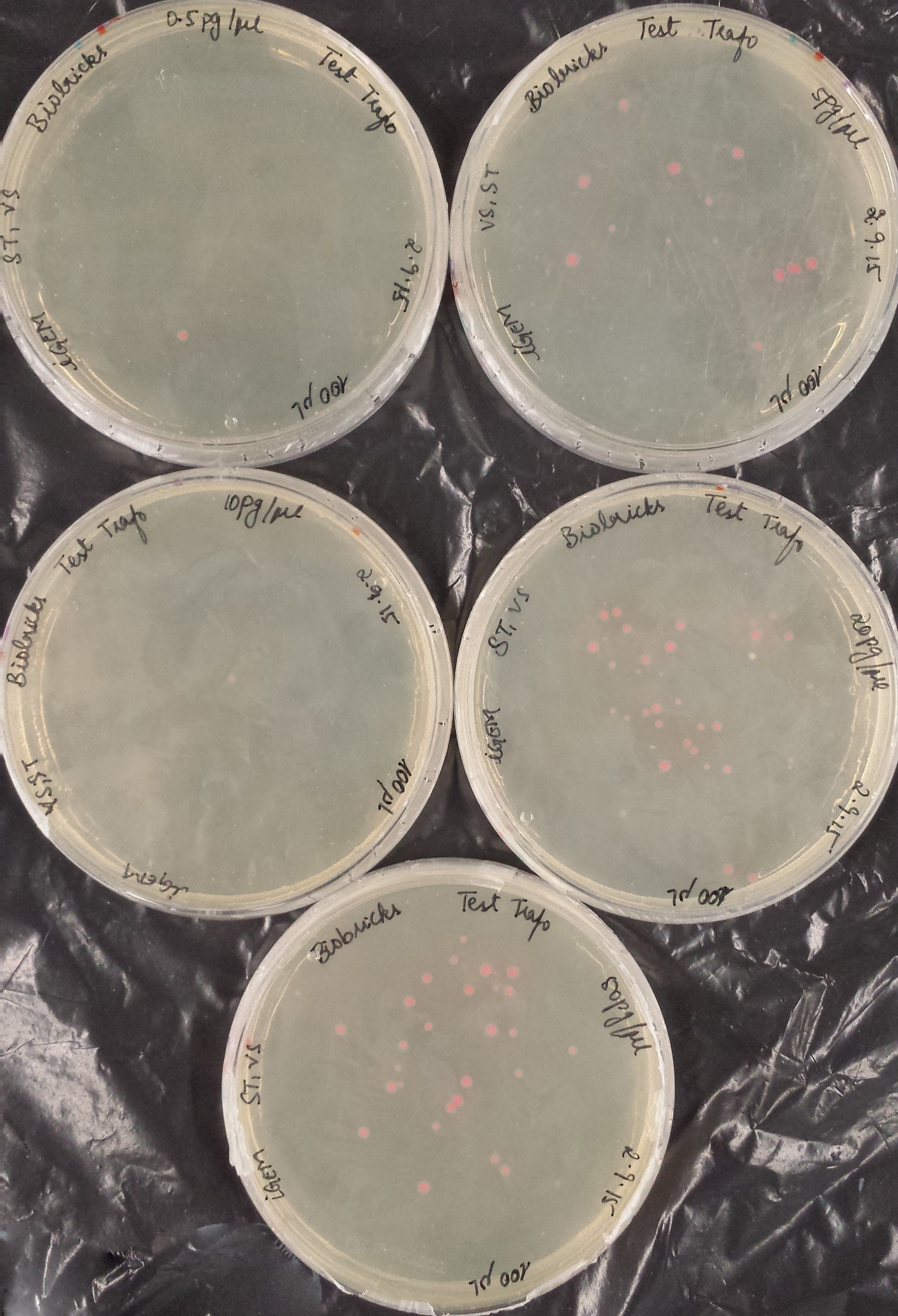
Before using our competent E. coli TOP10 cells in the important experiments, we used the Transformation Efficiency Kit to test the efficiency of our competent cells!
The kit includes five vials of each different DNA concentration: 50pg/μl, 20pg/μl, 10pg/μl, 5pg/μl, 0.5pg/μl of purified DNA from BBa_J04450 (RFP construct) in plasmid backbone pSB1C3.
The first test transformation (50 μl of competent cells, 20 μl plated) showed very poor results:
|
DNA concentration |
0.5pg/μl |
5pg/μl |
10pg/μl |
20pg/μl |
50pg/μl |
|
# of colonies |
0 |
0 |
2 |
0 |
13 |
|
efficiency (cfu) |
0 |
0 |
3.8x106 |
0 |
3.4x106 |
|
Results |
efficiency |
|
average |
1.4x106 |
|
weighted |
3.5x106 |
The second test transformation (200 μl of competent cells, 100 μl plated) showed still poor results but we decided to continue working with our cells:
|
DNA concentration |
0.5pg/μl |
5pg/μl |
10pg/μl |
20pg/μl |
50pg/μl |
|
# of colonies |
1 |
13 |
1 |
35 |
28 |
|
efficiency (cfu) |
8.0x106 |
1.0x107 |
4.0x105 |
7.0x106 |
2.3x106 |
|
Results |
efficiency |
|
average |
5.6x106 |
|
weighted |
3.7x106 |
RFP
RFP
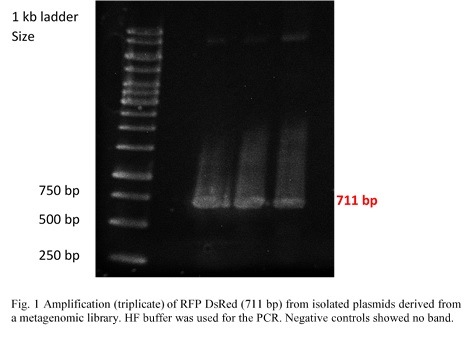
RFP (RFP DsRed) was amplified from pHT315_rfp by PCR (Fig.1). Colonies on a plate were given to us by the Applied and Genomic Microbiology department. Primers contained restriction sites for KpnI and SacI in order to make them compatible for insertion into the multiple cloning site of the pBAD/His A vector. The PCR products were examined in a 0,8% agarose gel by electrophoresis.
After purification of the PCR product it was ligated into pJET1.2 by subcloning (blunt end ligation). This vector serves to clone the fluorescent protein without triggering its activity, which may interact with the expression vector or the chosen E.coli strain.
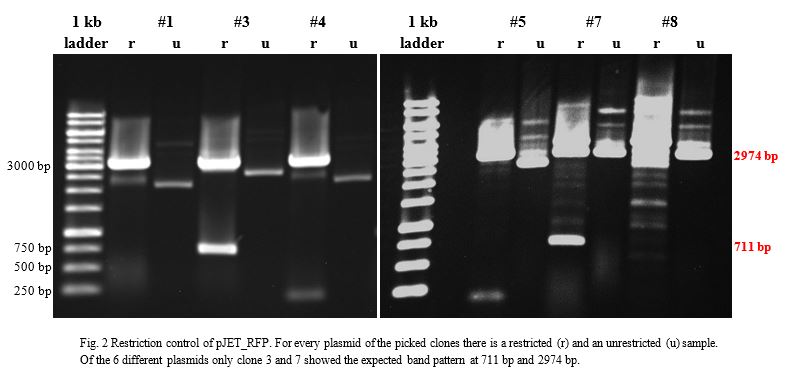
Ligation into pJET1.2 was followed according to the protocol in the methods collection. After over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit, restricted with KpnI and SacI and examined in a gel (Fig.2).
Sequencing of both samples was correct and proved that RFP_3 and RFP_7 were properly inserted into the pJET vector. We decided to keep on working with pJET_RFP_3. The plasmid was transformed with E.coli TOP10 and E.coli BL21. Cryocultures were frozen for a strain collection.
The next step was a big restriction of the whole plasmid extract with the correspondent pair of restriction enzymes. That allowed us to isolate the RFP inserts carrying the desired restriction sites (KpnI and SacI). The fragments were purified with the PEQLAB peqGOLD Gel Extraction Kit before T4 ligation. The RFP_3 insert was ligated into pBAD/His A by the T4 ligation system (sticky end ligation) according to the protocol in the methods collection and transformed into E.coli TOP10. This vector serves to express the fluorescent protein by triggering its activity.
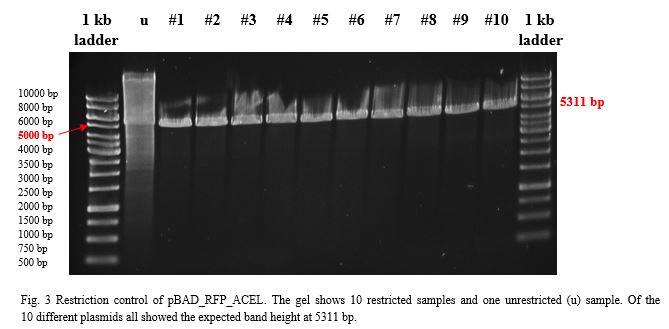
After over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit and restricted with KpnI and SacI.To build a component for our Flexosome, we decided to fuse pBAD_RFP with the ACEL (Acetivibrio cellulolyticus) dockerin. Both components, restricted with KpnI and SacI and purified with the PEQLAB peqGOLD Gel Extraction Kit were ligated by the T4 ligation system. After transformation into E.coli TOP10 and over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit, restricted with EcoRI and examined in a gel (Fig.3).
RESULTS
Sanger sequencing showed correct sequences for the Flexosome bit pBAD_RFP_ACEL.
MICROSCOPY
To check the activity of RFP, an induction series with 0,2%, 2% and 5% L-Arabinose in the cell culture medium was prepared. Fluorescence microscopy was performed with the RFP DsRed filter (excitation at 536 nm, emission at 582 nm). Unfortunately no fluorescence could be detected throughout the whole project with different constructs. Neither in the pJET_RFP_3, nor in the pBAD constructs (pBAD_RFP_3 and pBAD_RFP_ACEL) induced with 0,2%, 2% and 5% L-Arabinose in the medium. Nevertheless every time the DNA sequences were correct (Sanger sequencing). It might be a problem of expression.
FUTURE PLANS
Esterase and Phosphatase
Phosphatase and Esterase
Esterase was amplified from pET101_E064 and phosphatase from pTOPOXL_PLP07 by PCR for each enzyme (Fig.1). Both plasmids were given to us by the Applied and Genomic Microbiology department. Primers contained restriction sites KpnI and SacI for esterase and XhoI and PstI for the phosphatase in order to make them compatible for insertion into the multiple cloning site of the pBAD vector. The genes for these two enzymes had been found in screenings of metagenomic libraries.
After purification of the PCR products, they were ligated into pJET by blunt end ligation. This vector serves to clone the enzymes without triggering their activity, which may interact with the vector or the E.coli. In the case of phosphatase, the PCR products needed to be purified by gel extraction, to eliminate left over template plasmids, which could interfere with the transformation.
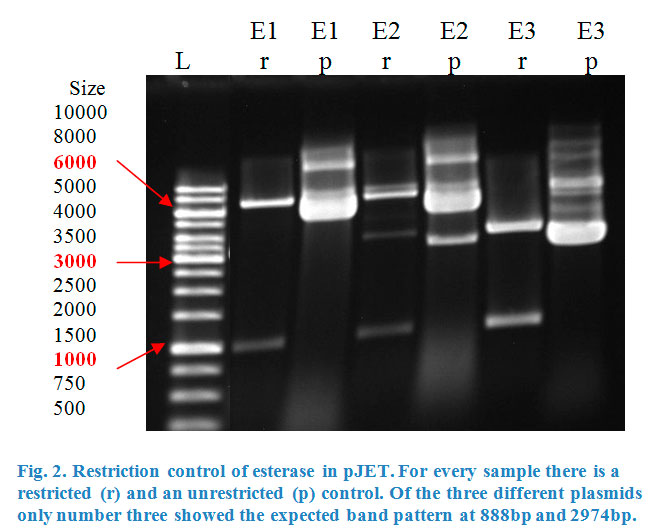
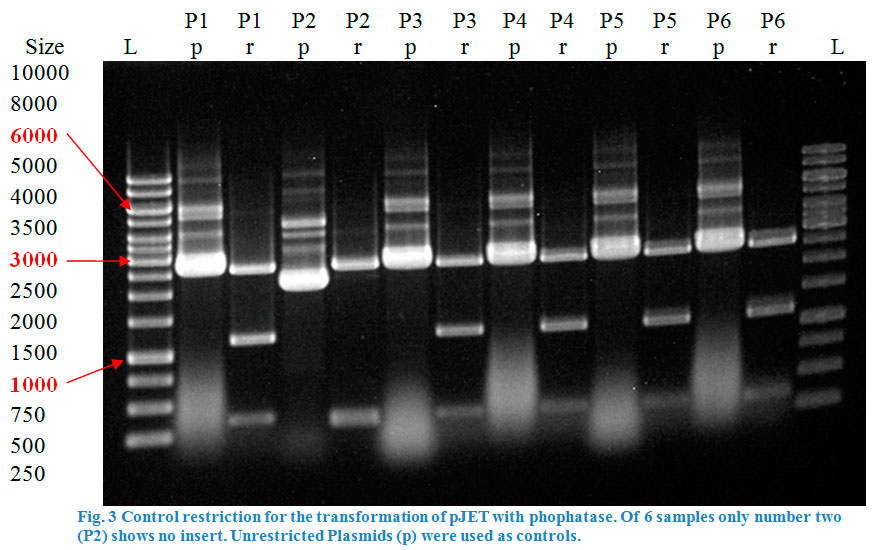
Ligation into pJET was followed according to the protocol in the methods collection. After over-night incubation colonies were picked, plasmids extracted with a Quiaprep spin Miniprep kit and restricted with either KpnI and SacI or XhoI and PstI, depending on the Enzyme (Fig.2&3).
Once restrictions showed the correct bands, both pJET containing esterase and phosphatase were sent for sequencing by the G2L laboratory.
The sequencing was successful and proved that both enzymes were properly inserted into the vector. The next step was a big restriction of the whole plasmid extract with the correspondent pair of restriction enzymes. That allowed us to isolate the enzyme inserts carrying the desired restriction sites. The fragments were purified with the peqGOLD Gel extraction kit.
The esterase was then ligated into the pBAD vector containing the CCEL (Clostridium cellulolyticum) dockering. Competent E. coli Top10 cells were transformed and the pBAD_Est_CCEL construct was isolated from the positive clones with a Quiaprep spin Miniprep kit.
A restriction control with SacI was run in a 0,8% agarose gel (Fig. 4). Samples #2, #3 and #9 correspond to the linear plasmid containing the insert.
The phosphatase fragment was fused to the pBAD and CTHE (Clostridium thermocellum) dockerin with the same strategy.
Project Achievements
What should this page contain?
Here you can describe the results of your project and your future plans.
- Clearly and objectively describe the results of your work.
- Future plans for the project
- Considerations for replicating the experiments
You can also include a list of bullet points (and links) of the successes and failures you have had over your summer. It is a quick reference page for the judges to see what you achieved during your summer.
- A list of linked bullet points of the successful results during your project
- A list of linked bullet points of the unsuccessful results during your project. This is about being scientifically honest. If you worked on an area for a long time with no success, tell us so we know where you put your effort.
Inspiration
See how other teams presented their results.