Difference between revisions of "Team:Goettingen/Results"
| Line 599: | Line 599: | ||
<p> | <p> | ||
</html> | </html> | ||
| − | [[ | + | [[File:IGEM-presentation_final_team_Goettingen.ppt]] |
<html> | <html> | ||
</p> | </p> | ||
<p> | <p> | ||
</html> | </html> | ||
| − | [[ | + | [[File:Poster_teamGoettingen_2015.pdf]] |
<html> | <html> | ||
</p> | </p> | ||
</div> | </div> | ||
</html> | </html> | ||
Revision as of 21:59, 20 October 2015
Project Results
Transformation Efficiency Kit, RFP construct (iGEM)
Before using our competent E. coli TOP10 cells in the important experiments, we used the Transformation Efficiency Kit to test the efficiency of our competent cells!
The kit includes five vials of each different DNA concentration: 50 pg/μl, 20 pg/μl, 10 pg/μl, 5 pg/μl, 0.5 pg/μl of purified DNA from BBa_J04450 (RFP construct) in plasmid backbone pSB1C3.
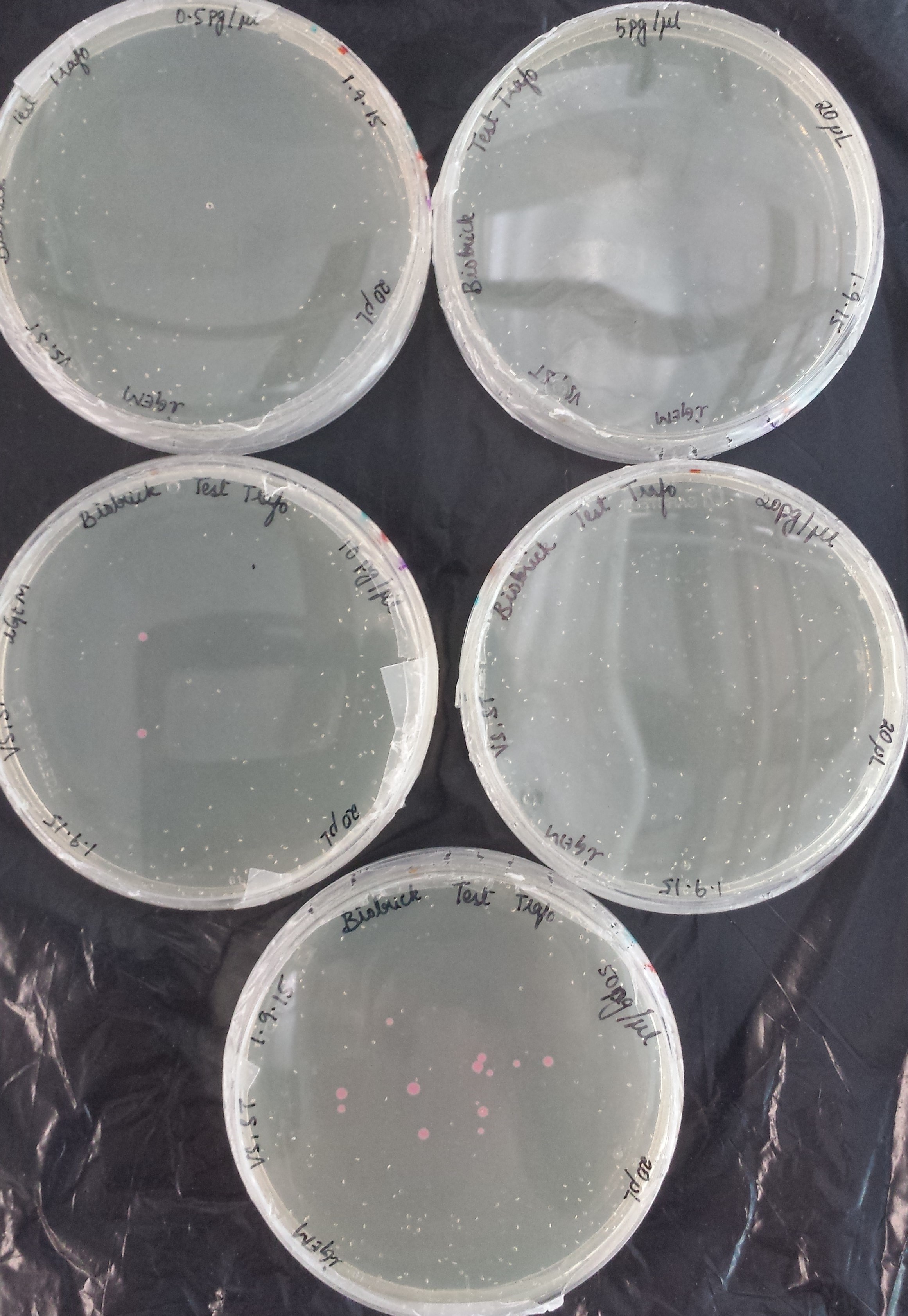
The first test transformation (50 μl of competent cells, 20 μl plated) showed very poor results:
|
DNA concentration |
0.5 pg/μl |
5 pg/μl |
10 pg/μl |
20 pg/μl |
50 pg/μl |
|
# of colonies |
0 |
0 |
2 |
0 |
13 |
|
efficiency (cfu) |
0 |
0 |
3.8x 106 |
0 |
3.4x 106 |
|
Results |
efficiency |
|
average |
1.4x 106 |
|
weighted |
3.5x 106 |
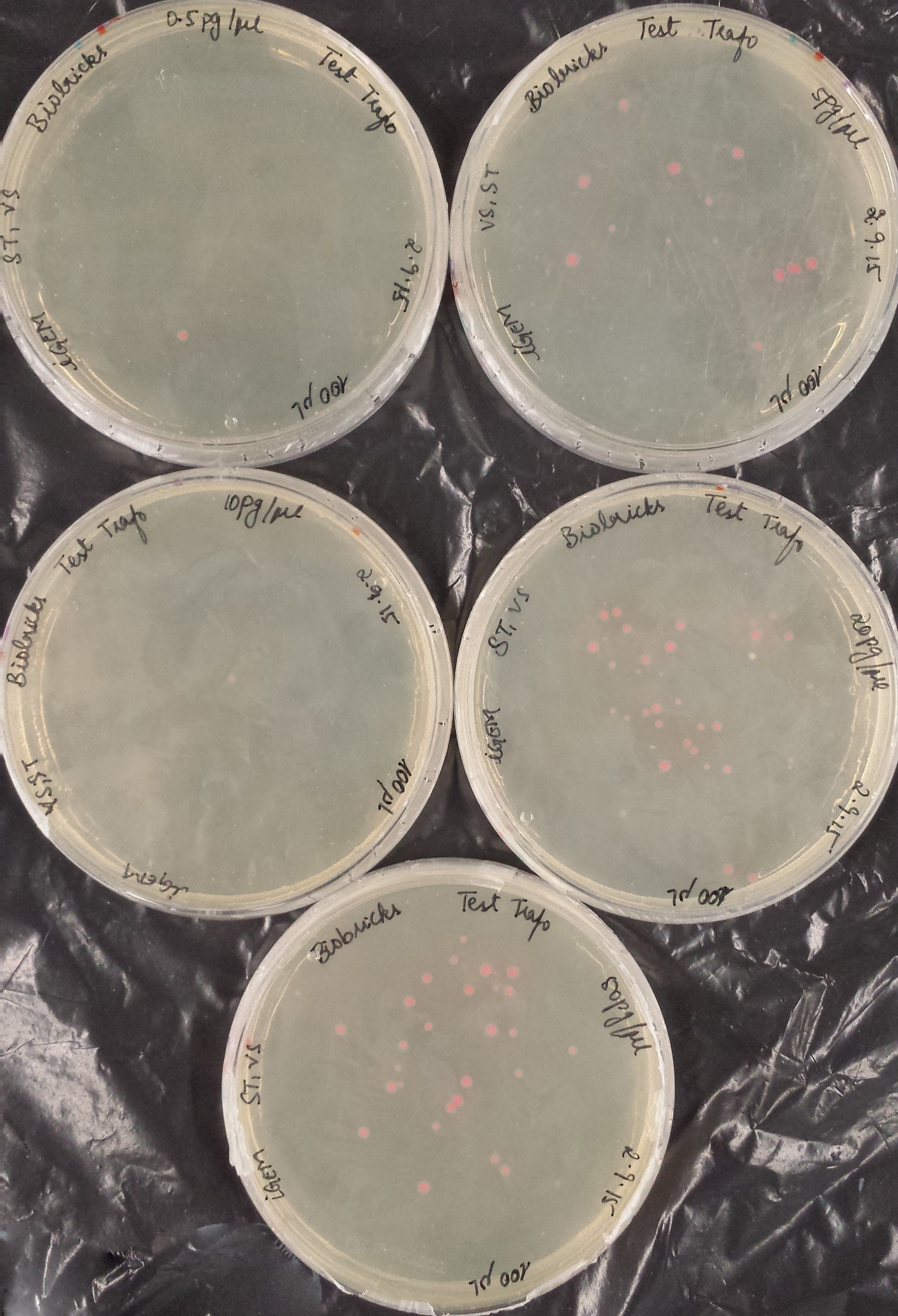
The second test transformation (200 μl of competent cells, 100 μl plated) showed still poor results but we decided to continue working with our cells:
|
DNA concentration |
0.5 pg/μl |
5 pg/μl |
10 pg/μl |
20 pg/μl |
50 pg/μl |
|
# of colonies |
1 |
13 |
1 |
35 |
28 |
|
efficiency (cfu) |
8.0x 106 |
1.0x 107 |
4.0x 105 |
7.0x 106 |
2.3x 106 |
|
Results |
efficiency |
|
average |
5.6x 106 |
|
weighted |
3.7x 106 |
RFP
RFP
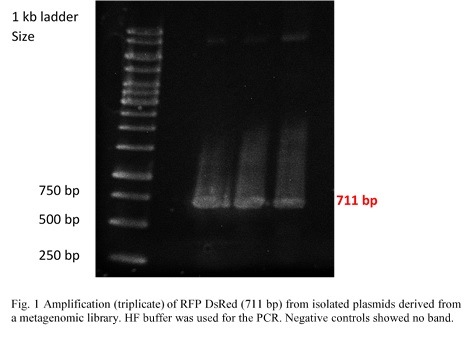
RFP (RFP DsRed) was amplified from pHT315_rfp by PCR (Fig.1). Colonies on a plate were given to us by the Applied and Genomic Microbiology department. Primers contained restriction sites for KpnI and SacI in order to make them compatible for insertion into the multiple cloning site of the pBAD/His A vector. The PCR products were examined in a 0,8% agarose gel by electrophoresis.
After purification of the PCR product it was ligated into pJET1.2 by subcloning (blunt end ligation). This vector serves to clone the fluorescent protein without triggering its activity, which may interact with the expression vector or the chosen E.coli strain.
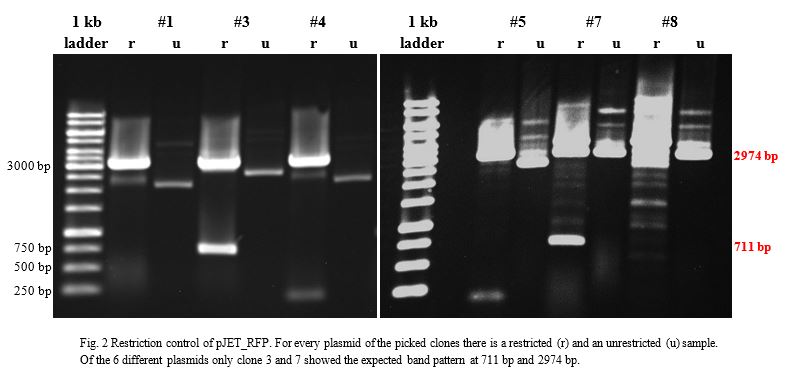
Ligation into pJET1.2 was followed according to the protocol in the methods collection. After over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit, restricted with KpnI and SacI and examined in a gel (Fig.2).
Sequencing of both samples was correct and proved that RFP_3 and RFP_7 were properly inserted into the pJET vector. We decided to keep on working with pJET_RFP_3. The plasmid was transformed with E.coli TOP10 and E.coli BL21. Cryocultures were frozen for a strain collection.
The next step was a big restriction of the whole plasmid extract with the correspondent pair of restriction enzymes. That allowed us to isolate the RFP inserts carrying the desired restriction sites (KpnI and SacI). The fragments were purified with the PEQLAB peqGOLD Gel Extraction Kit before T4 ligation. The RFP_3 insert was ligated into pBAD/His A by the T4 ligation system (sticky end ligation) according to the protocol in the methods collection and transformed into E.coli TOP10. This vector serves to express the fluorescent protein by triggering its activity.
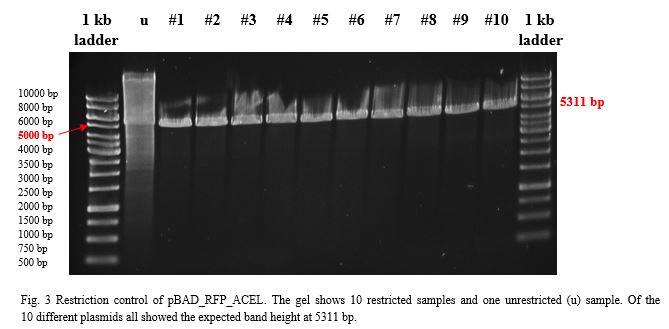
After over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit and restricted with KpnI and SacI.To build a component for our Flexosome, we decided to fuse pBAD_RFP with the ACEL (Acetivibrio cellulolyticus) dockerin. Both components, restricted with KpnI and SacI and purified with the PEQLAB peqGOLD Gel Extraction Kit were ligated by the T4 ligation system. After transformation into E.coli TOP10 and over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit, restricted with EcoRI and examined in a gel (Fig.3).
RESULTS
Sanger sequencing showed correct sequences for the Flexosome bit pBAD_RFP_ACEL.
MICROSCOPY
To check the activity of RFP, an induction series with 0,2%, 2% and 5% L-Arabinose in the cell culture medium was prepared. Fluorescence microscopy was performed with the RFP DsRed filter (excitation at 536 nm, emission at 582 nm). Unfortunately no fluorescence could be detected throughout the whole project with different constructs. Neither in the pJET_RFP_3, nor in the pBAD constructs (pBAD_RFP_3 and pBAD_RFP_ACEL) induced with 0,2%, 2% and 5% L-Arabinose in the medium. Since the DNA sequences have been confirmed to be correct by Sanger sequencing it might be a problem of expression.
FUTURE PLANS
Esterase and Phosphatase
Phosphatase and Esterase
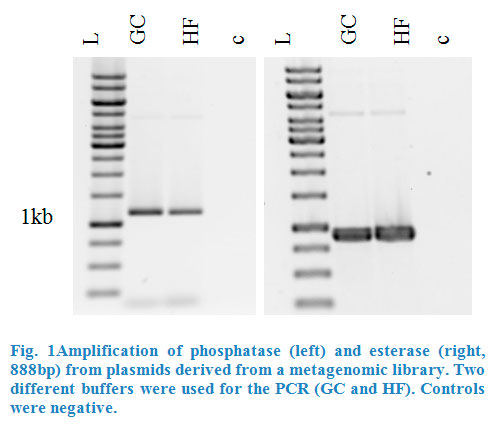
Esterase was amplified from pET101_E064 and phosphatase from pTOPOXL_PLP07 by PCR for each enzyme (Fig.1). Both plasmids were given to us by the Applied and Genomic Microbiology department. Primers contained restriction sites KpnI and SacI for esterase and XhoI and PstI for the phosphatase in order to make them compatible for insertion into the multiple cloning site of the pBAD vector. The genes for these two enzymes had been found in screenings of metagenomic libraries.
After purification of the PCR products, they were ligated into pJET by blunt end ligation. This vector serves to clone the enzymes without triggering their activity, which may interact with the vector or the E.coli. In the case of phosphatase, the PCR products needed to be purified by gel extraction, to eliminate left over template plasmids, which could interfere with the transformation.
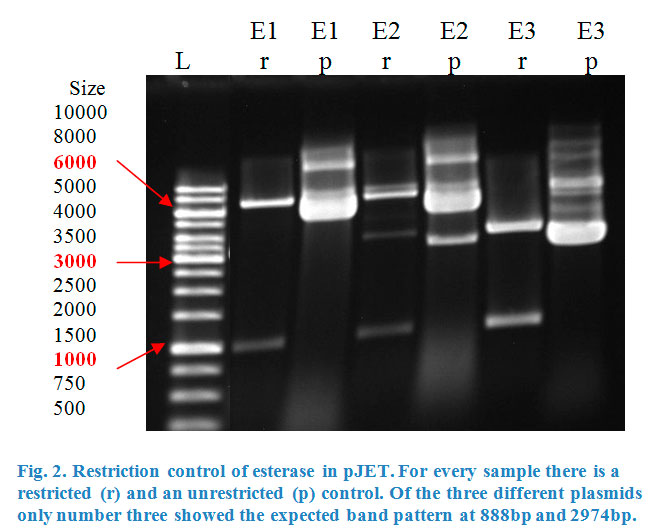
Ligation into pJET was followed according to the protocol in the methods collection. After over-night incubation colonies were picked, plasmids extracted with a Quiaprep spin Miniprep kit and restricted with either KpnI and SacI or XhoI and PstI, depending on the Enzyme (Fig.2&3).
Once restrictions showed the correct bands, both pJET containing esterase and phosphatase were sent for sequencing by the G2L laboratory.
The sequencing was successful and proved that both enzymes were properly inserted into the vector. The next step was a big restriction of the whole plasmid extract with the correspondent pair of restriction enzymes. That allowed us to isolate the enzyme inserts carrying the desired restriction sites. The fragments were purified with the peqGOLD Gel extraction kit.
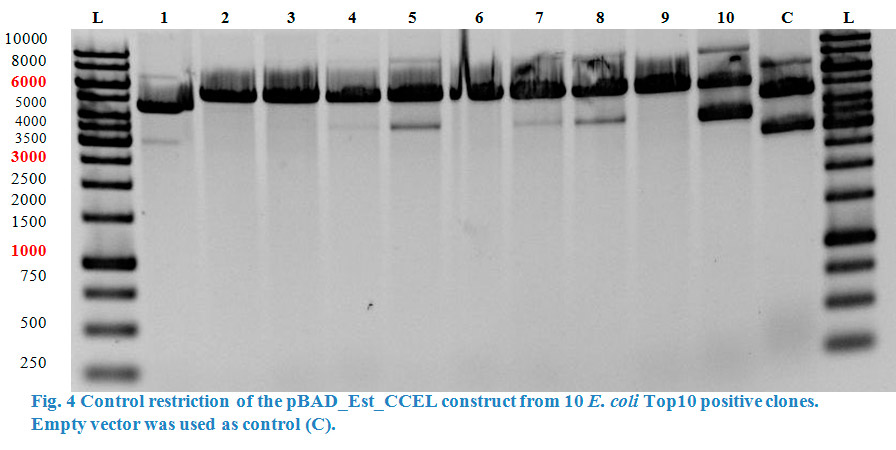
The esterase was then ligated into the pBAD vector containing the CCEL (Clostridium cellulolyticum) dockering. Competent E. coli Top10 cells were transformed and the pBAD_Est_CCEL construct was isolated from the positive clones with a Quiaprep spin Miniprep kit.
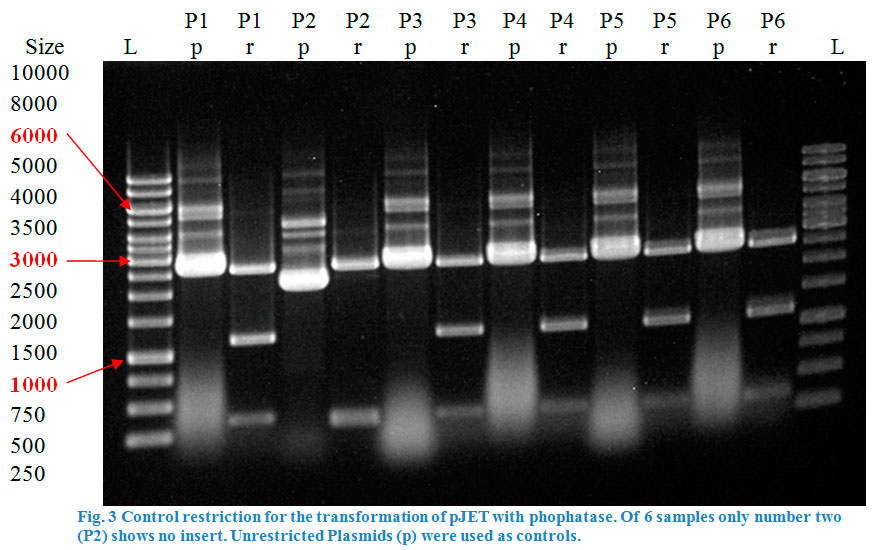
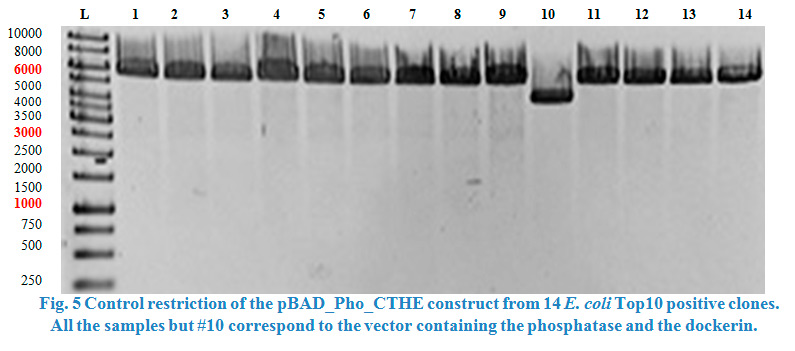
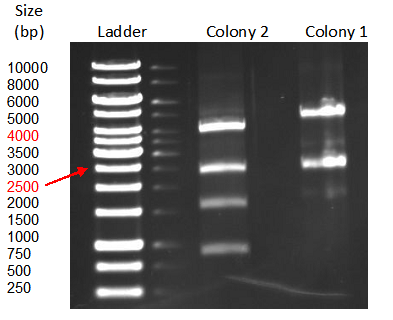
A restriction control with SacI for the esterase constructs was run in a 0,8% agarose gel (Fig. 4). Samples #2, #3 and #9 correspond to the linear plasmid containing the insert. Phosphatase construct was control restricted with EcoRI (Fig. 5), in this case 13 of 14 samples showed successful insertion of the enzyme fragment into the vector.
Future plans
The next steps include changing to a better expression vector in order to test and measure the activity of the enzymes, especially to ensure that the fusion with the dockerins is not interfering with the function of the enzymes. Finally the aim is to combine the purified enzyme-dockerin proteins with the synthetic scaffold and prove the self assemby of the Flexosome tool.
Cellulase
The cellulase gene was amplified using PCR. The primers which were used contained the restriction sites for ligation into the pBAD vector. The cellulase PCR products were then checked on 0.8 % agarose gel.
The cellulase PCR products were then purified by using the QIAquick® PCR Purification Kit (QIAGEN). After the PCR products were purified, the concentration of DNA was measured by Nanodrop 2000c Spectrophotometer.
Restriction control of pBAD_CCEL and cellulase PCR product
The purified cellulase PCR product and the pBAD_CCEL vector were restricted by following a double digest protocol with SacI and PvuII enzymes. After restriction control products were purified by using the QIAquick® PCR Purification Kit (QIAGEN) the concentration of DNA was measured by Nanodrop 2000c Spectrophotometer.
Ligation of Cellulase into pBAD- CCEL
The cellulase gene was then ligated into the pBAD vector containing the CCEL (Clostridium
cellulolyticum ) dockerin.
For ligation of the cellulase PCR product into the restricted pBAD_CCEL vector, the Thermo Fisher T4 Ligase Sticky End Ligation protocol was followed.
Transformation of pBAD_CellulaseCCEL into chemically competent Top10 E. coli cells
pBAD_CellulaseCCEL was transformed into competent Top10 E. coli cells by using the heat shock transformation method.
Colonies from transformation plates were inoculated into LB medium containing ampicillin (100 µg/ml).
Plasmid Extraction: peqGold Plasmid Miniprep Kit
Plasmid extraction was done for colonies inoculated in LB medium by using peqGold Plasmid Miniprep kit and plasmid concentration was measured by using Nanodrop 2000c Spectrophotometer.
Restriction control was done with the enzymes SacI and PvuII from Thermo fisher scientific. Restricted plasmids were checked on a 0.8 % agarose gel.
The correct insert was confirmed through sequencing.
BioBricks
Dockerin Biobricks
The dockerins ACEL (Acetivibrio cellulolyticus), BCEL (Bacteroides cellulosolvens), CCEL (Clostridium cellulolyticum) and CTHE (Clostridium thermocellum) were amplified from pBAD_ACEL, pBAD_BCEL, pBAD_CCEL and pBAD_CTHE with primers to add the desired EcoRI and PstI restriction sites by PCR. Primers contained restriction sites for EcoRI and PstI in order to make them compatible for insertion into the iGEM shipping vector pSB1C3.
After purification, the PCR products were restricted with EcoRI and PstI, as well as pSB1C3. Afterwards the restricted dockerins were ligated into pSB1C3 by T4 ligation (sticky end ligation) and transformed into E. coli TOP10.
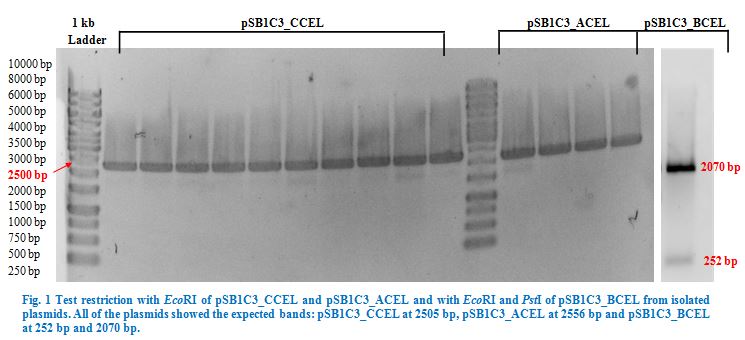
Ligation into pSB1C3 was followed according to the protocol in the methods collection. After over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit and test restricted with EcoRI (pSB1C3_CCEL and pSB1C3_ACEL) and EcoRI and PstI (pSB1C3_BCEL) (Fig.1). We lost CTHE due to failing transformations at this point.
Once restriction controls showed the correct bands, pSB1C3_CCEL, pSB1C3_ACEL and pSB1C3_BCEL were sent for sequencing.
Colour BioBricks
We also wanted to improve already existing BioBricks by fusing our dockerins to the colours eforRed (BBa_K592012) and amilCP (BBa_K592009) that were submitted from the University of Uppsala (Sweden) in 2011.
So we chose to fuse eforRed to BCEL and amilCP to CCEL. Since the plasmids of the two colour proteins were not distributed with the current plate of the iGEM Distribution Kit, we ordered those parts including the right restriction sites as gBlocks from IDT.
As a first step eforRed and amilCP were restricted with KpnI and SacI to make them compatible with our dockerins in pBAD.
After purification restricted colours were ligated into pBAD_BCEL and pBAD_CCEL by T4 ligation (sticky end ligation) and transformed into E. coli TOP10.
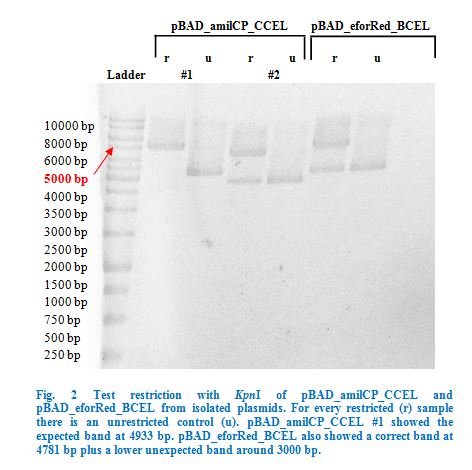
Ligation into pBAD was followed according to the protocol in the methods collection. After over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit and test restricted with KpnI (Fig.2).
Once restriction controls showed the correct bands, pBAD_amilCP_CCEL #1 and pBAD_eforRed_BCEL were sent for sequencing. Sequencing showed that both plasmids were correct.
To make our constructs compatible with the iGEM shipping vector pSB1C3 the desired EcoRI and PstI restrictions sites were added by PCR.
After purification, the PCR products were restricted with EcoRI and PstI, the same with pSB1C3, and afterwards ligated into pSB1C3 by T4 ligation (sticky end ligation) and transformed into E. coli TOP10.
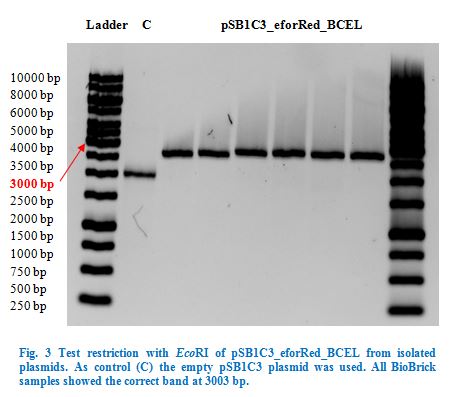
After over-night incubation colonies were picked, plasmids extracted with the QIAGEN QIAprep Spin Miniprep Kit and test restricted with EcoRI (Fig.3). Due to the reason that our dockerin CCEL contains an internal PstI restriction site we continued working only with the eforRed_BCEL construct.
Once restriction controls showed the correct band, pSB1C3_eforRed_BCEL was sent for sequencing. Sequencing showed that the plasmids were correct.
RESULTS
Sequencing showed that the dockerins ACEL and CCEL contain an internal PstI restriction site. Therefore all the constructs containing ACEL or CCEL showed in the end truncated dockerin sequences and could not be send in as BioBricks.
We also build a construct where the enzyme esterase was fused to our CCEL dockerin (pBAD_Est_CCEL) following the same strategy. But again, due to the reason that our dockerin CCEL contains an internal PstI restriction site we had to stop our work at this point.
Furthermore we collaborated with the current iGEM team of Aachen and tried to fuse three of their enzymes to our constructs but could not finish our work here.
Sequences of pSB1C3_BCEL and pSB1C3_eforRed_BCEL were correct and submitted as BioBricks to iGEM:
pSB1C3_BCEL (BBa_K1865000)
pSB1C3_eforRed_BCEL (BBa_K1865001)
Scaffoldin Purification
Scaffoldin purification
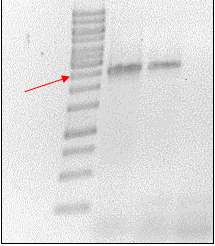
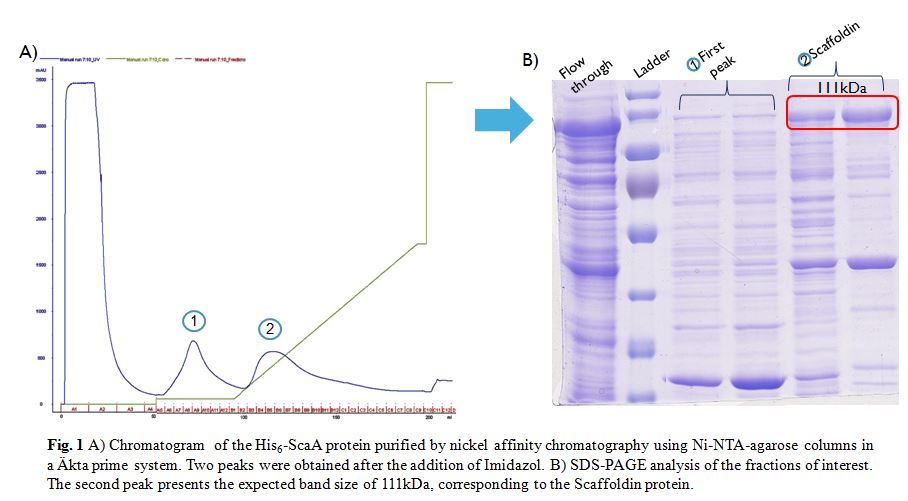
After induction according to the methods collection, the targeted ScaA scaffoldin was purified by nickel affinity chromatography using Ni-NTA-agarose columns in a Äkta prime system (Fig. 1A). Following the addition of Imidazol two peaks were obtained. The fractions of interest were visualized in a SDS-PAGE gel (Fig. 1B). The gel showed that only the second fraction presents the expected size of 111kDa. However some other bands appeared along this fraction, therefore in order to obtain even more pure protein a Size Exclusion Chromatography (SEC) was done.
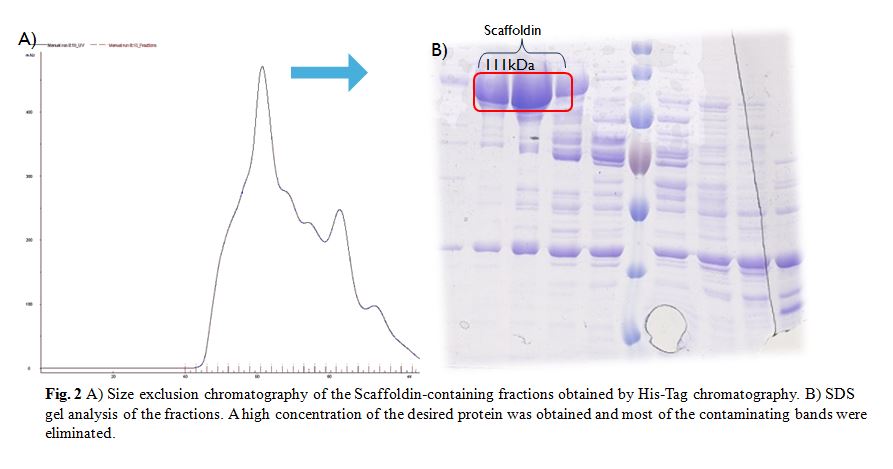
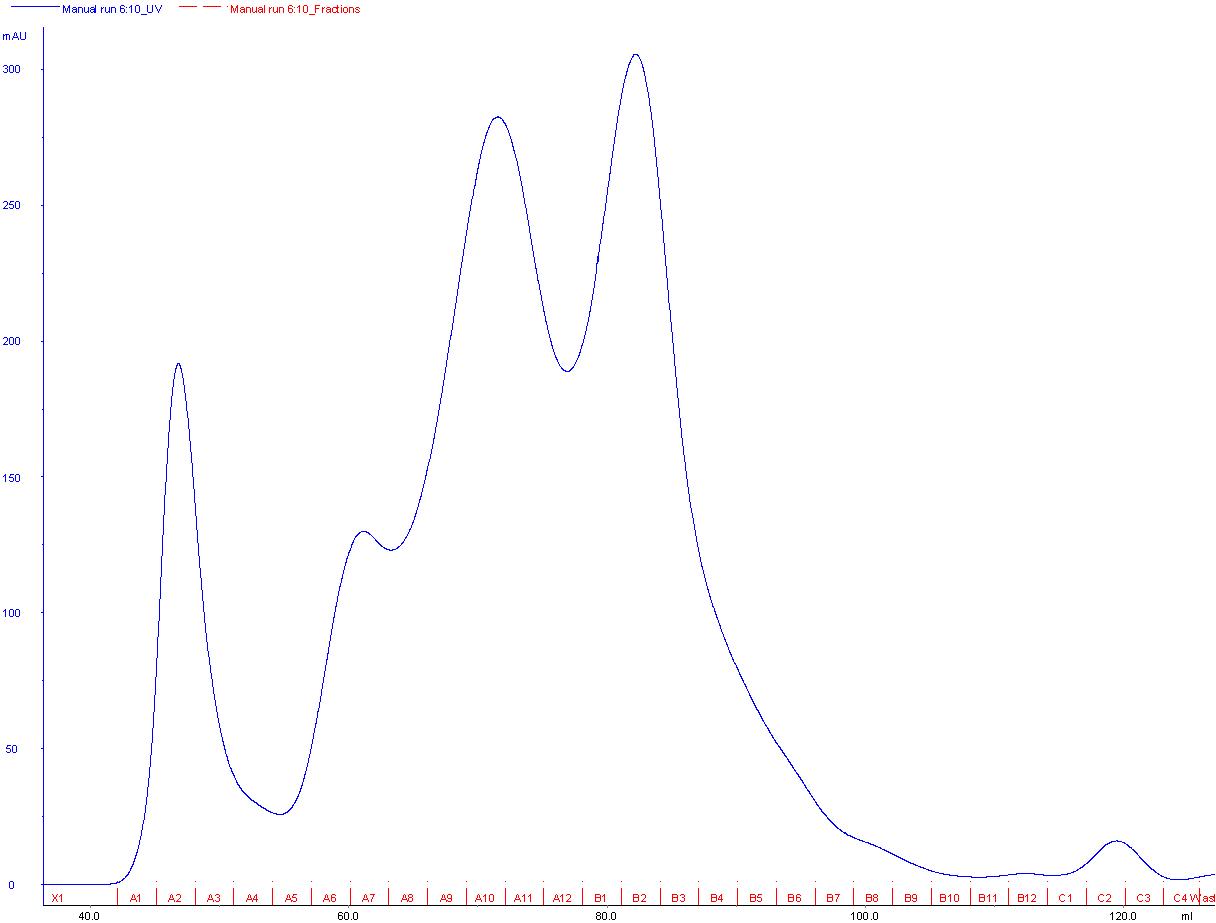
The SEC was done with a HiLoad Superdex 200 16/600 column (GE Healthcare) (Fig. 2A). SDS gel shows that a high concentration of the desired protein was obtained and most of the previous contaminating bands were eliminated. Nevertheless there was an unknown band of low weight present in every fraction, apparently it could be a second protein from E. coli with some affinity to the Scaffoldin than is then is co-purified with it (Fig. 2B).
Future plans include to incubate the Scafoldin with the purified enzyme-dockerin construct in order to prove that the Flexosome can assembly by itself.
Copurification
Copurification of Scaffoldin and Enzyme-Dockerin
The main peaks after gelfiltration from the scaffoldin CTHE (see results scaffoldin) and CellulaseCCEL (Fig.1) were pooled and concentrated.
They were then mixed and incubated together at 37°C for 15 min with agitation.
The solution was then used in another gelfiltration (Fig.2). The resulting peaks were run on a SDS-gel (Fig.3) and checked for cellulase activity.
The first peak from the copurification showed a distinct cellulase activity. As the peak ran considerably higher than expected than either the scaffoldin or the CellulaseCCEL in the gelfitration, it can be expected that this peak contains a successfully formed complex between CellulaseCCEL and the CTHE scaffoldin.
Giant Jamboree: Poster and Presentation
Did you missed our presentation? Do you want to see our poster again? No problem, here you can find the files: