Team:Czech Republic/Design
Design and Functional Prototype
The IOD band is a project built on system level integration. At the outset, it seemed like an impossible goal for a single iGEM project especially as there are no precedents for many of the system parts. So the idea was stripped to its brass tacks to uncover three fundamental modules that prove the diagnostic and engineering principles in a tractable manner.
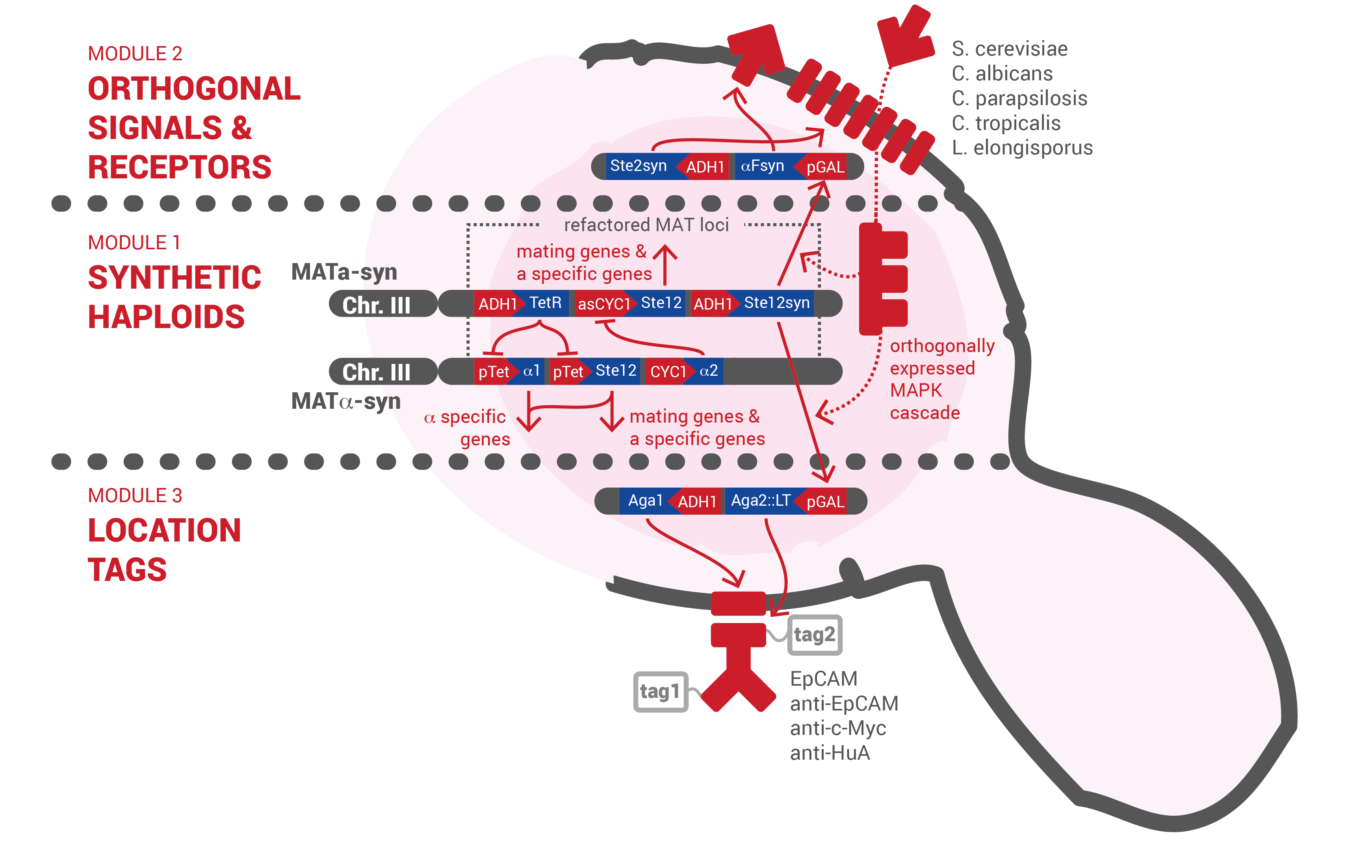
Module 1 builds synthetic haploid strains with refactored mating loci that are conjugated to make a functional IOD. These strains have the wild-type mating phenotype and differentially express a reprogrammed signalling pathway in their diploid state proving the feasibility of the clone-free assembly concept.
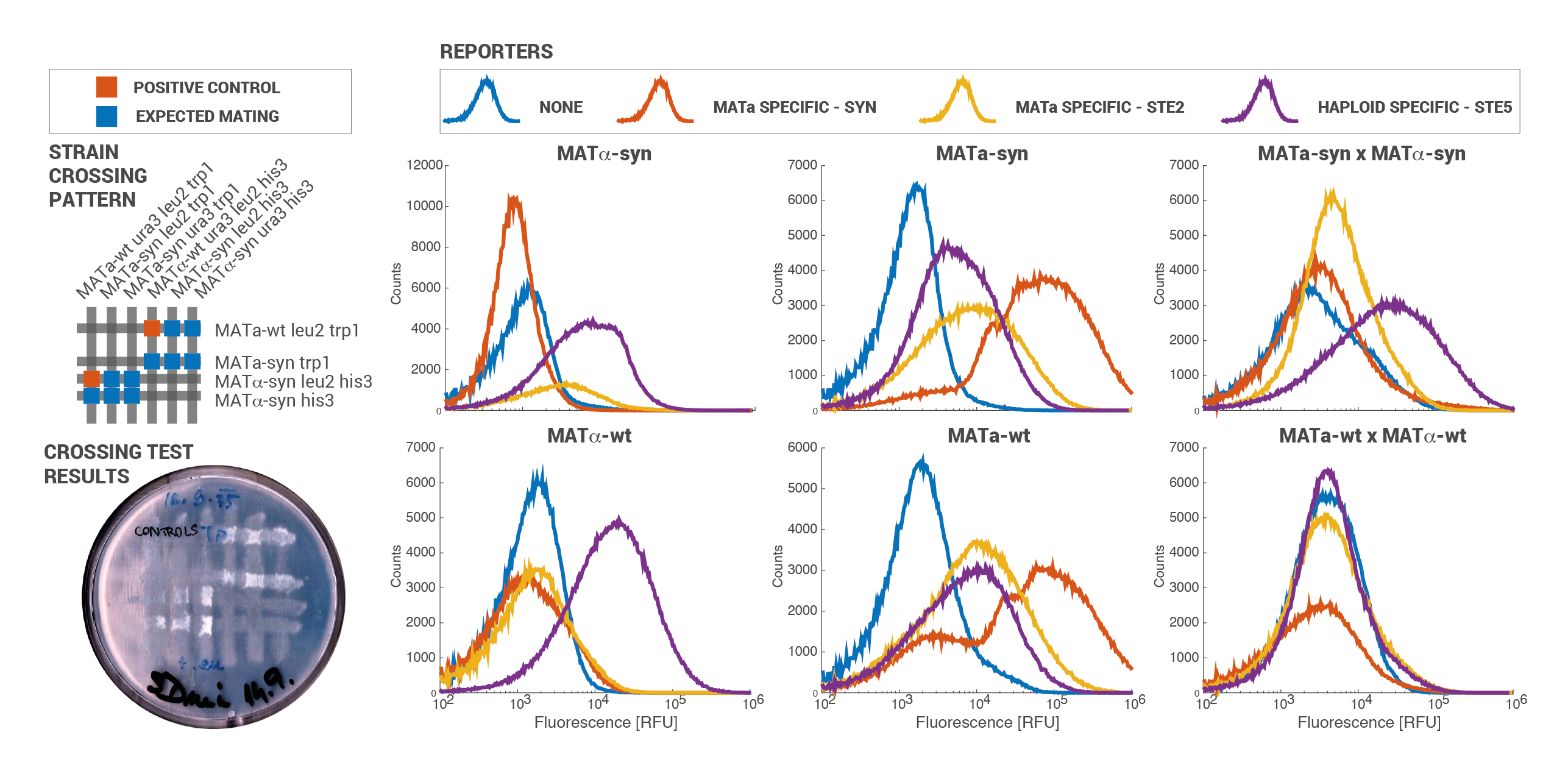
Functional prototypes of both MATa and MATx synthetic haploid strains were constructed and experimentally validated. The successful construction and integration of the refactored loci was confirmed using traditional methods and proves the effectivity of the flanking homologous regions. The retained wild-type mating phenotype of both synthetic haploid strains was validated by direct mating tests where strains were crossed and plated onto minimal medium. This shows that the synthetic strains are easily conjugated. Differential expression of haploid-specific and mating-type specific genes with respect to the wild-type strains was validated by flow cytometry. Using the developed collection of reporter plasmids, it was shown that mating-type specific genes are expressed only in their respective haploid strains and that haploid-specific genes are expressed in both the haploid and diploid strains. This demonstrates that the signaling cascade can be used orthogonally in haploid strains for conjugation purposes and in diploid strains for IOD intercellular signal detection. For the first time in iGEM, the function of a tunable a-specific promoter (a hybrid of the CYC1 and a-specific operon domains) was also validated.
Module 2 builds a set of orthogonal pheromones and receptors. These pheromone-receptor pairs enable specific localized signalling proving the feasibility of multichannel signal transmission underlying logic operations necessary for reliable diagnosis.
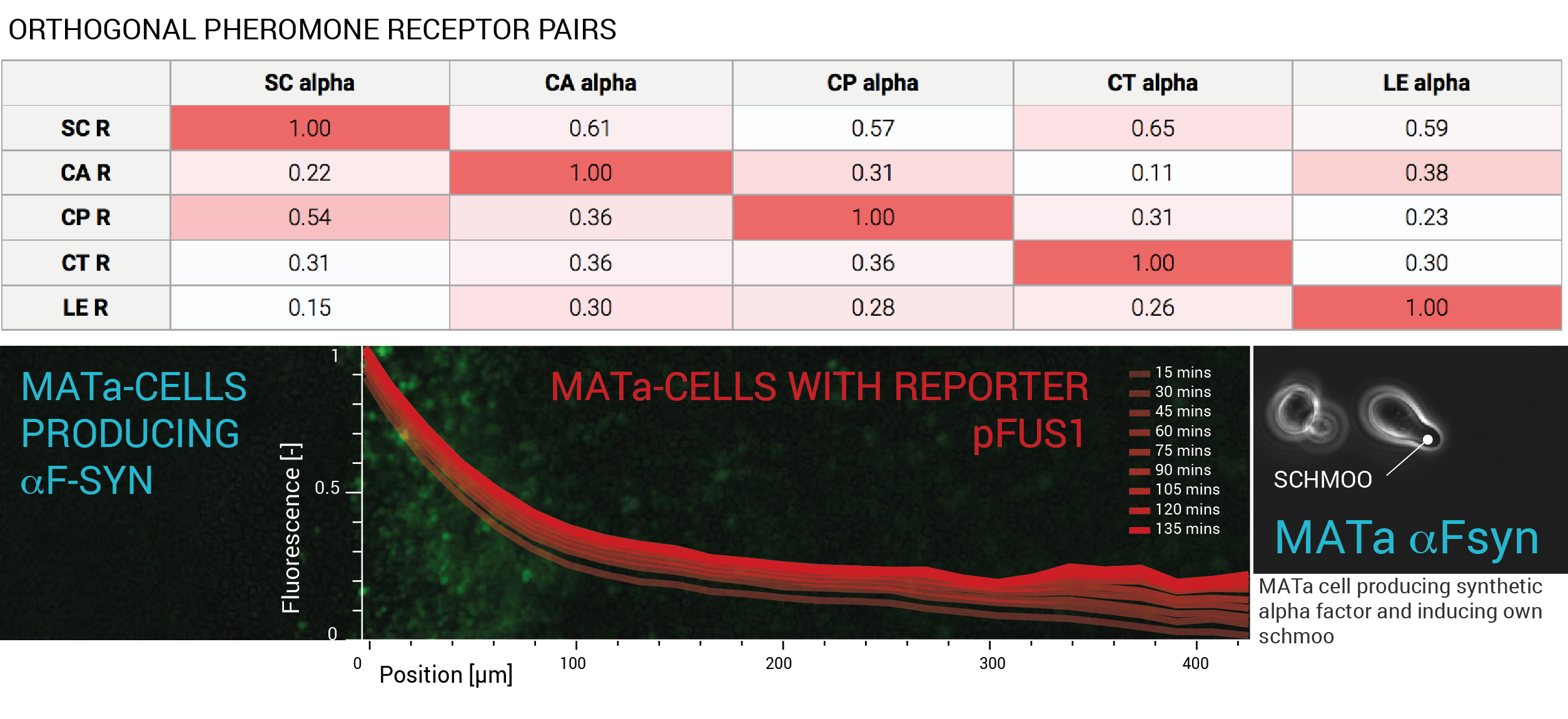
For the first time, orthogonality of five different yeast pheromone-receptor pairs was validated. Alpha factor from various yeast strains was synthetically produced in S. cerevisiae and used to induce the pheromone pathway rewired with the corresponding Ste2 receptors. Fluorescence measurements from the pheromone pathway reporter pFUS1 showed maximal activation in matching pairs that significantly dominated any cross talk between the non-matching pairs. Furthermore, functionality of the synthetic sequences was extensively tested using both physiological and molecular assays. To prove the designed alpha factor, which was truncated to exclude repeats induces mating, a MATa cell synthetically expressing the alpha phermone was shown to schmoo after induction. To prove localization of pheromone based signalling to the immediate neighbourhood of the transmitting cell, a microfluidic experiment was designed that directly measured the signalling range in saturated cultures. An extrapolation of the results indicates that in a culture with OD600 greater than 3.0 only cells in direct contact will transmit signal.
Module 3 builds a set of location tags that recognize common tumor surface markers and agglutinate cell populations. Location tags displayed in the correct conformation strengthen cell-cell interactions to enable localization of signal transmission.
For the first time, an antibody (anti-EpCam), the most commonly used CTC marker (EpCAM), and the human antigen A antibody (anti-HuA) were displayed on the surface of the cell wall. Successful construction of the display plasmids was confirmed using traditional methods. Successful epitope export and display were then validated by immunofluorescent staining using an upstream hemagglutinin tag. Clumping of cells with complimentary location tags (i.e., anti-EpCAM and Ep-CAM) was shown qualitatively using microscopy and by visual inspection of stained slides with mounted cultures. While all cells have a tendency to aggregate that varies with incubation period and temperature, cells displaying complimentary location tags always generated visibly larger aggregates than their controls. Functioning of location tags in plasma, which is the natural medium for the IOD band diagnostic, was further proven by cell clumping consistently observed in mixed samples of yeast cells with the anti-HuA location tag and red blood cells from a type A donor. The control samples, where anti-HuA location tag display was not induced, showed negligible aggregation. These experiments also prove the compatibility of yeast cells with blood samples as no significant clumping or other abnormalities that would indicate overall concept infeasibility were observed in control samples.

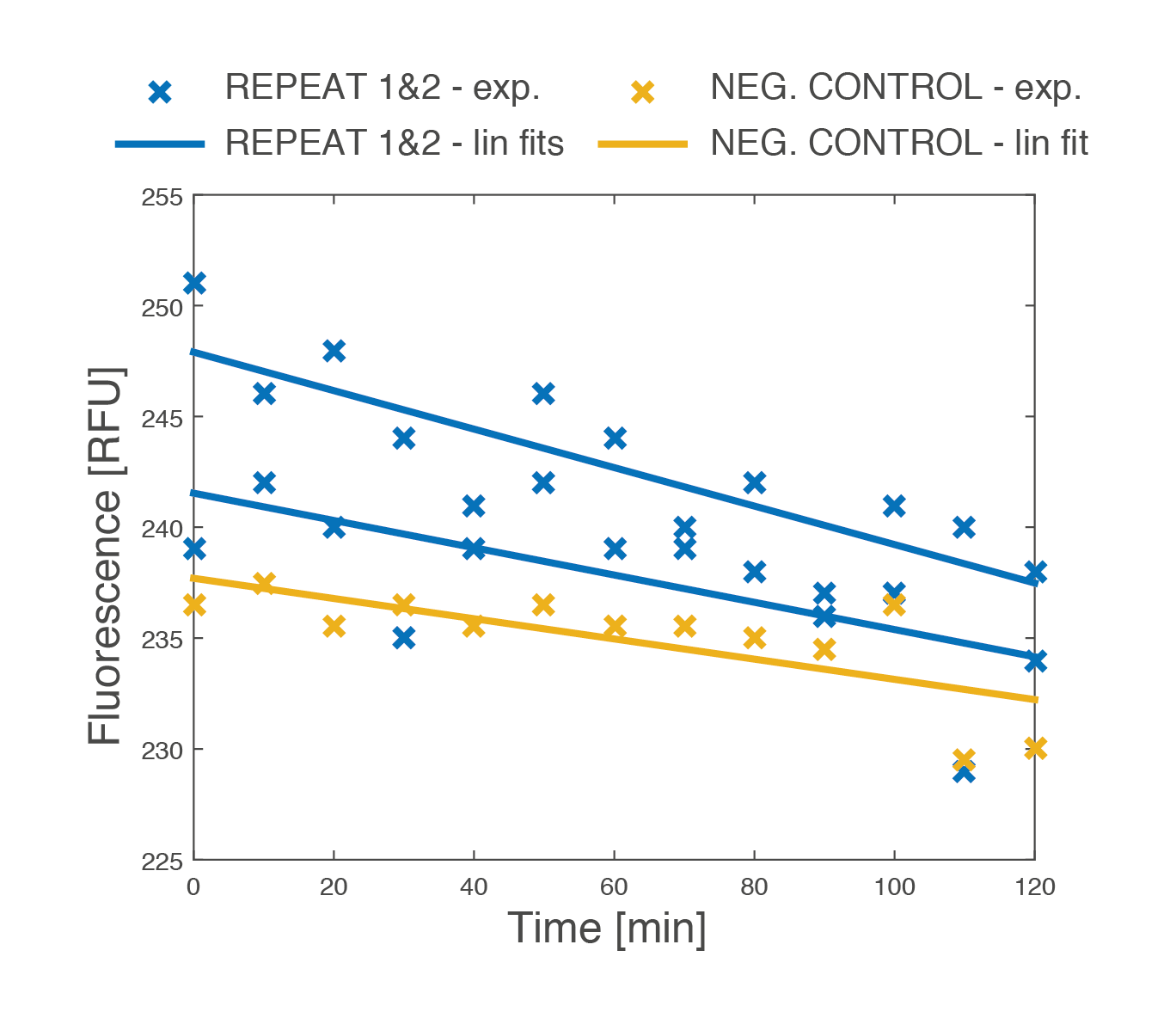
A final master experiment integrated the location tag and signalling functions. Signal transmission enabled by location tag display was shown in two strains constitutively displaying complimentary tags (EpCAM and anti-EpCAM). Strain A represented an IOD bound to a CTC: a transformed MATa strain displaying a fragment of the common epithelial CTC marker EpCAM and carrying a pheromone response reporter (GFP upstream of pFUS1). Strain B represented a second IOD that recognises the EpCAM marker and activates the IOD band response: a transformed MATa strain synthetically producing the S. cerevisiae pheromone. Localised signal transmission is difficult to quantify. Qualitatively fluorescence microscopy revealed greater intensity levels in observed aggregates. Microplate assays suggest modest activation of Strain A in mixed cultures when compared to background activation in pure cultures and controls. Mixed culture controls with strains lacking the location tags exhibited no activation.
It is important to note that modeling and microfluidics were extensively used to support the experimental work. Modeling proved robustness of simple signal transmission networks in a simulation setting with naturally occurring signaling noise. In-house replicated microfluidic chips provided the precise environment necessary to quantify the individual cell-cell interactions. Without these engineering tools, realization and validation of such a complex project would have been as impossible as the IOD band idea seemed at the outset.