Team:UCLA/Notebook/Honeybee Silk/3 August 2015
Analyzing SDS PAGE Gel results
- Danny ran the SDS PAGE gel for the honeybee purification protocol on 7/30 and Megan stained the gel for visualization of 7/31.
- The gel was run, and then stored overnight in destain buffer due to logistical reasons before being stained the next day.
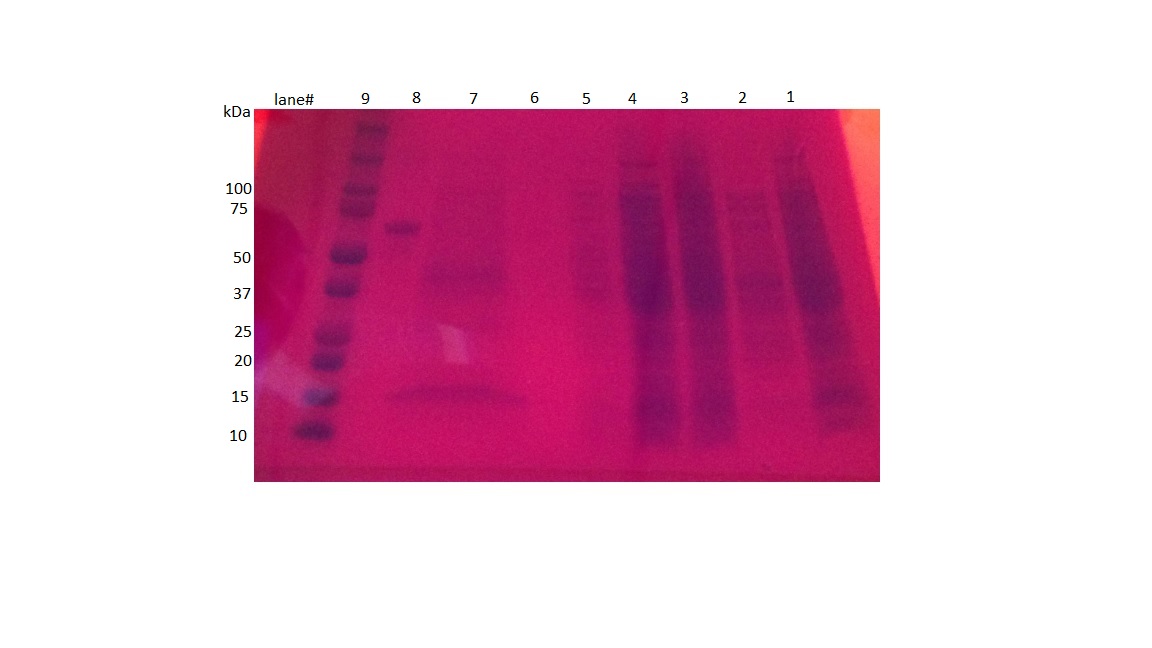
- Here is the gel
- Refer to 7/29 for more detailed description of when each fraction was taken.
| F1 (lane 1) | Full cell lysate using BugBuster lysis system |
|---|---|
| S1 (lane 2) | Supernatant after spinning down full cell lysate |
| F2 (lane 3) | Full cell lysate #2 after resuspending pellet in Bugbuster |
| F3 (lane 4) | Full cell lysate #3 after adding lysozyme and DNase |
| S2 (lane 5) | Supernatant after first wash step. |
| S3 (lane 6) | Supernatant after second wash step |
| Final product (lane 7) | Purified inclusion body in SDS |
| positive control (lane 8) | BSA positive control |
| ladder (lane 9) | BioRad dual color ladder |
Analysis
- This gel illuminates some things about our expression and purification procedure.
- The band at around 40 kDa matches the expected size for honeybee silk based off of the paper we are using as a guideline, so we are getting at least some of the expected product.
- There is a large contaminating band at 15 kDa. It is presumably an insoluble protein, just likely the honeybee inclusion bodies.
- Because there seems to be almost no protein in the S3 fraction, it would seem that the second wash step is unnecessary and should be skipped next time in order to increase yield.