Difference between revisions of "Team:Vilnius-Lithuania/Labjournal"
| (27 intermediate revisions by 4 users not shown) | |||
| Line 3: | Line 3: | ||
{{Team:Vilnius-Lithuania/Vilnius15_Socials}} | {{Team:Vilnius-Lithuania/Vilnius15_Socials}} | ||
| + | <html> | ||
<div id="wikicontent-container"> | <div id="wikicontent-container"> | ||
<div id="wikicontent"> | <div id="wikicontent"> | ||
| − | |||
<body> | <body> | ||
<style> | <style> | ||
.panel-heading [data-toggle="collapse"]:after { | .panel-heading [data-toggle="collapse"]:after { | ||
| + | display: inline-block; | ||
font-family: 'Glyphicons Halflings'; | font-family: 'Glyphicons Halflings'; | ||
| − | content: "\ | + | /*content: "\00ab"; "play" icon */ |
| + | content: url(https://static.igem.org/mediawiki/2015/d/d8/Vilnius15_chevron_up_h2.png); | ||
position: absolute; | position: absolute; | ||
| − | top: | + | top: 5px; |
| − | right: | + | right: 5px; |
color: #DE564B; | color: #DE564B; | ||
font-size: 18px; | font-size: 18px; | ||
line-height: 22px; | line-height: 22px; | ||
/* rotate "play" icon from > (right arrow) to down arrow */ | /* rotate "play" icon from > (right arrow) to down arrow */ | ||
| − | -webkit-transform: rotate( | + | -webkit-transform: rotate(180deg); |
| − | -moz-transform: rotate( | + | -moz-transform: rotate(180deg); |
| − | -ms-transform: rotate( | + | -ms-transform: rotate(180deg); |
| − | -o-transform: rotate( | + | -o-transform: rotate(180deg); |
| − | transform: rotate( | + | transform: rotate(180deg); |
} | } | ||
.panel-heading [data-toggle="collapse"].collapsed:after { | .panel-heading [data-toggle="collapse"].collapsed:after { | ||
| + | display: inline-block; | ||
| + | content: url(https://static.igem.org/mediawiki/2015/e/e1/Vilnius15_chevron_up.png); | ||
/* rotate "play" icon from > (right arrow) to ^ (up arrow) */ | /* rotate "play" icon from > (right arrow) to ^ (up arrow) */ | ||
| − | -webkit-transform: rotate( | + | -webkit-transform: rotate(0deg); |
| − | -moz-transform: rotate( | + | -moz-transform: rotate(0deg); |
| − | -ms-transform: rotate( | + | -ms-transform: rotate(0deg); |
| − | -o-transform: rotate( | + | -o-transform: rotate(0deg); |
| − | transform: rotate( | + | transform: rotate(0deg); |
color: #454444; | color: #454444; | ||
} | } | ||
| + | |||
| + | .panel-heading a.accordion-toggle { | ||
| + | color: rgb(62,62,62); | ||
| + | text-decoration: none; | ||
| + | } | ||
</style> | </style> | ||
<section class="container-fluid" style="padding: 0px; padding-top: 20px;"> | <section class="container-fluid" style="padding: 0px; padding-top: 20px;"> | ||
| − | |||
| − | |||
| Line 47: | Line 54: | ||
<div class="panel-title"> | <div class="panel-title"> | ||
<h3 style="padding: 0px; margin: 0px;">Parts construction</h3> | <h3 style="padding: 0px; margin: 0px;">Parts construction</h3> | ||
| + | |||
<div class="row"> | <div class="row"> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
| Line 53: | Line 61: | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
Written by: <strong>IR</strong><br /> | Written by: <strong>IR</strong><br /> | ||
| − | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>MB</strong>, <strong>IS</strong> | + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>MB</strong>, <strong>IS</strong> |
| − | + | </div> | |
</div> | </div> | ||
</div> | </div> | ||
| Line 98: | Line 106: | ||
Extraction from gel<br/> | Extraction from gel<br/> | ||
L10 and L11 are produced.<br/> | L10 and L11 are produced.<br/> | ||
| − | Conc.: L11 – 2,4 ng/ | + | Conc.: L11 – 2,4 ng/μL. |
</td> | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>07 13</td> | ||
| + | <td>P10</td> | ||
| + | <td> | ||
| + | Repeated restriction of P10 with EcoRI and XBal (+FastAp).<br /> | ||
| + | Conc.: L10 – 45,3 ng/μL. L10 and L11 ligation.<br /> | ||
| + | P14 is produced.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>07 14</td> | ||
| + | <td>Bacteria with P14</td> | ||
| + | <td>Colony PCR of P14 bacteria. 1<sup>st</sup> and 2<sup>nd</sup> colonies are inoculated.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td></td> | ||
| + | <td>P14.2</td> | ||
| + | <td>Sequencing</td> | ||
</tr> | </tr> | ||
</tbody> | </tbody> | ||
| Line 112: | Line 138: | ||
</div> | </div> | ||
</div> | </div> | ||
| + | |||
| + | </html> | ||
| + | [[File:Vilnius15_P14_2_protokolas.png|500px|thumb|<strong>Figure 1:</strong> Restriction analysis of P14. Restriction fragments after digestion with EcoRI and SpeI. M – O’Gene ruler DNA ladder mix, K – undigested plasmid, E+S – plasmid digested with EcoRI and SpeI.|left]] | ||
| + | <html> | ||
</div> | </div> | ||
| Line 126: | Line 156: | ||
<div class="row"> | <div class="row"> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
| − | Part: <strong>P15.1</strong> ( | + | Part: <strong>P15.1</strong> (BBa_K1773003) |
</div> | </div> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
Written by: <strong>IR</strong><br /> | Written by: <strong>IR</strong><br /> | ||
| − | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>MB</strong>, <strong>IS</strong> | + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>MB</strong>, <strong>IS</strong> |
</div> | </div> | ||
</div> | </div> | ||
| Line 141: | Line 171: | ||
<div class="col-md-12"> | <div class="col-md-12"> | ||
<h3><strong>About</strong></h3> | <h3><strong>About</strong></h3> | ||
| − | <p>Cas3 with mutated restriction sites<br />L7+L15. Concentration - 261 ng/μL</p> | + | <p>Cas3 with mutated restriction sites.<br />L7+L15. Concentration - 261 ng/μL</p> |
</div> | </div> | ||
</div> | </div> | ||
| Line 244: | Line 274: | ||
</div> | </div> | ||
</div> | </div> | ||
| + | </html> | ||
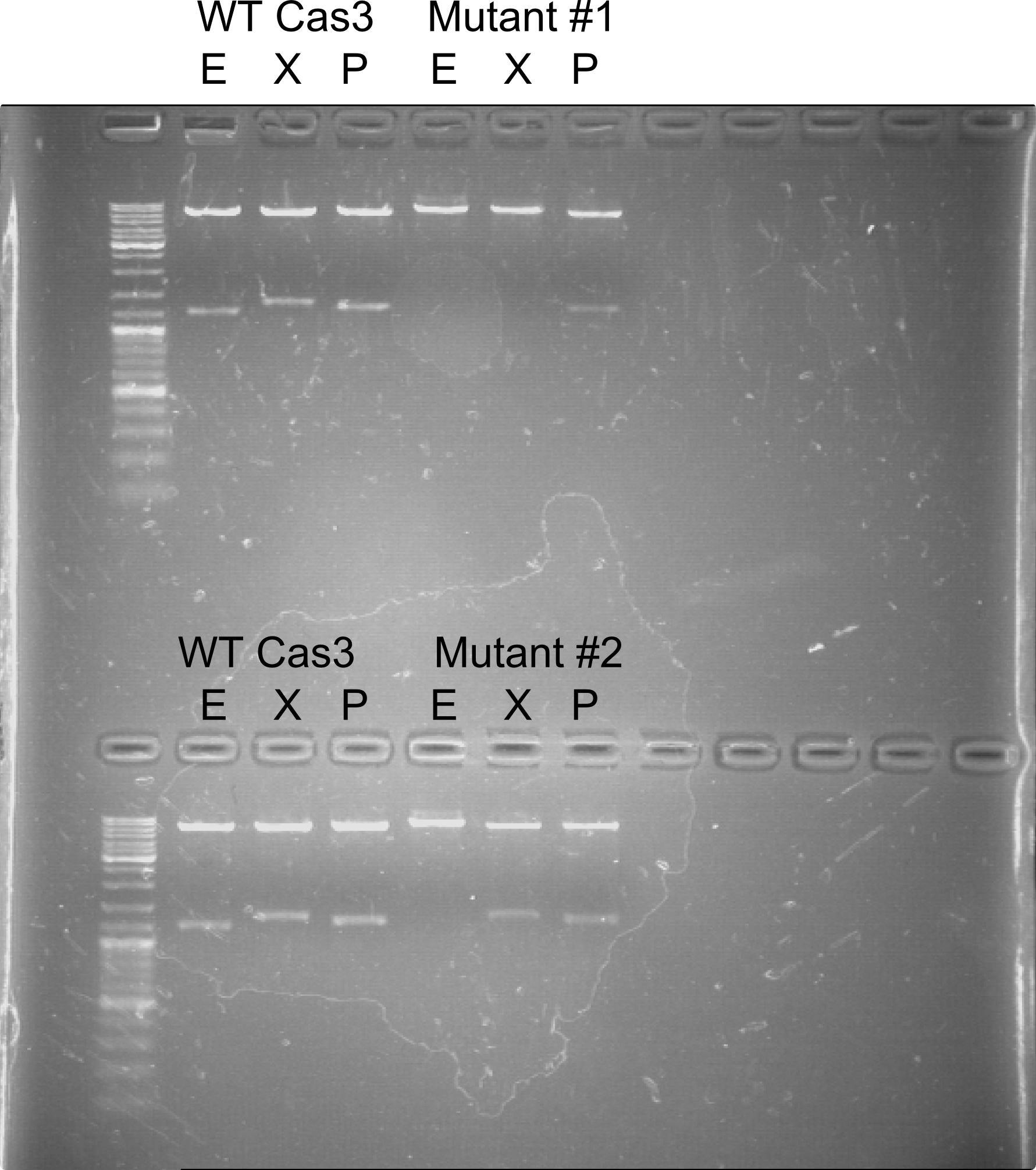
| + | [[File:Vilnius15_P15_1_construction_protokolas_1.png|500px|thumb|<strong>Figure 1:</strong> First restriction analysis. Restriction with EcoRI (E), XbaI (X) and PstI (P) of Wild type (WT) Cas3 gene compared to mutated Cas3 genes. The first mutant plasmid has two successful mutations, whereas the second mutant plasmid only has one successfully mutated restriction site.|left]] | ||
| + | <html> | ||
| + | |||
| + | </html> | ||
| + | [[File:Vilnius15_P15_1_construction_protokolas_2.png|500px|thumb|<strong>Figure 2:</strong> Restriction analysis of Cas3 mutagenesis second run. K - undigested Wild type Cas3 plasmid. E - digested with EcoRI; X - digested with XbaI; P - digested with PstI.|left]] | ||
| + | <html> | ||
</div> | </div> | ||
| Line 258: | Line 295: | ||
<div class="row"> | <div class="row"> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
| − | Parts: <strong>P26 | + | Parts: |
| + | <ul style="list-style: none; padding: 0px; margin: 0px;"> | ||
| + | <li><strong>P26</strong> (BBa_K1773016)</li> | ||
| + | <li><strong>P27</strong> (BBa_K1773017)</li> | ||
| + | <li><strong>P28</strong> (BBa_K1773018)</li> | ||
| + | <li><strong>P29</strong> (BBa_K1773012)</li> | ||
| + | </ul> | ||
</div> | </div> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
Written by: <strong>IR</strong><br /> | Written by: <strong>IR</strong><br /> | ||
| − | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>BB</strong>, <strong>IS</strong> | + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>BB</strong>, <strong>IS</strong> |
</div> | </div> | ||
</div> | </div> | ||
| Line 273: | Line 316: | ||
<div class="col-md-12"> | <div class="col-md-12"> | ||
<h3><strong>About</strong></h3> | <h3><strong>About</strong></h3> | ||
| − | <p>P26 – L7+L26 pLux/cI+SRBS<br /> | + | <p> |
| + | P26 – L7+L26 pLux/cI+SRBS<br /> | ||
P27 – L7+L27 pLux/cI+MRBS <br /> | P27 – L7+L27 pLux/cI+MRBS <br /> | ||
P28 – L7+L28 pLux/cI+WRBS <br /> | P28 – L7+L28 pLux/cI+WRBS <br /> | ||
| Line 293: | Line 337: | ||
<tr> | <tr> | ||
<td>07 27</td> | <td>07 27</td> | ||
| − | <td>ST9<br /> | + | <td> |
| + | ST9<br /> | ||
ST10<br /> | ST10<br /> | ||
ST11<br /> | ST11<br /> | ||
| Line 314: | Line 359: | ||
<tr> | <tr> | ||
<td>07 28</td> | <td>07 28</td> | ||
| − | <td>L7<br /> | + | <td> |
| + | L7<br /> | ||
L26<br /> | L26<br /> | ||
L27<br /> | L27<br /> | ||
| Line 325: | Line 371: | ||
<td>07 29 - <br /> - 08 05</td> | <td>07 29 - <br /> - 08 05</td> | ||
<td></td> | <td></td> | ||
| − | <td>L7+L26 colony PCR with ST15 and VR.<br /> | + | <td> |
| + | L7+L26 colony PCR with ST15 and VR.<br /> | ||
L7+L27 colony PCR with ST15 and VR.<br /> | L7+L27 colony PCR with ST15 and VR.<br /> | ||
L7+L28 colony PCR with ST15 and VR.<br /> | L7+L28 colony PCR with ST15 and VR.<br /> | ||
| Line 363: | Line 410: | ||
<div class="row"> | <div class="row"> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
| − | Part: <strong>P30</strong> ( | + | Part: <strong>P30</strong> (BBa_K1773007) |
</div> | </div> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
Written by: <strong>IR</strong><br /> | Written by: <strong>IR</strong><br /> | ||
| − | Performed by: <strong>ŠT</strong> | + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>IS</strong> |
</div> | </div> | ||
</div> | </div> | ||
| Line 469: | Line 516: | ||
<div class="row"> | <div class="row"> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
| − | + | Parts: | |
| − | + | <ul style="list-style: none; padding: 0px; margin: 0px;"> | |
| + | <li><strong>P35</strong> (BBa_K1773019)</li> | ||
| + | <li><strong>P36</strong> (BBa_K1773020)</li> | ||
| + | <li><strong>P37</strong> (BBa_K1773021)</li> | ||
| + | </ul> | ||
| + | </div> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
Written by: <strong>IR</strong><br /> | Written by: <strong>IR</strong><br /> | ||
| − | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>BB</strong>, <strong>IS</strong> | + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>BB</strong>, <strong>IS</strong> |
</div> | </div> | ||
</div> | </div> | ||
| Line 537: | Line 589: | ||
L7+L28 colony PCR with ST15 and VR<br /> | L7+L28 colony PCR with ST15 and VR<br /> | ||
1st and 2nd colonies are inoculated.<br /> | 1st and 2nd colonies are inoculated.<br /> | ||
| − | P35, P36 and | + | P35, P36 and P37 are produced<br /> |
P35, P36, P37 restriction with NspI.</td> | P35, P36, P37 restriction with NspI.</td> | ||
</tr> | </tr> | ||
| Line 571: | Line 623: | ||
<div class="row"> | <div class="row"> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
| − | Part: <strong>Cd1</strong> ( | + | Part: <strong>Cd1</strong> (BBa_K1773005) |
</div> | </div> | ||
<div class="col-md-6"> | <div class="col-md-6"> | ||
Written by: <strong>IR</strong><br /> | Written by: <strong>IR</strong><br /> | ||
| − | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>MB</strong>, <strong>IS</strong>, <strong>IO</strong> | + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>MB</strong>, <strong>IS</strong>, <strong>IO</strong> |
</div> | </div> | ||
</div> | </div> | ||
| Line 670: | Line 722: | ||
</div> | </div> | ||
</div> | </div> | ||
| + | |||
| + | </html> | ||
| + | [[File:Vilnius15_Parts_Cd1_protokolas.png|500px|thumb|<strong>Figure 1:</strong> . Cascade restriction analysis. After mutant digestion with EcoRI no DNA linearisation was present, compared to the non-mutated Cascade control (K).|left]] | ||
| + | <html> | ||
| + | |||
</div> | </div> | ||
| Line 676: | Line 733: | ||
<!-- /MethodBox --> | <!-- /MethodBox --> | ||
| + | <!-- MethodBox --> | ||
| + | <div class="panel panel-default" style="position: relative"> | ||
| + | <div class="panel-heading"> | ||
| + | <a class="accordion-toggle collapsed" data-toggle="collapse" data-parent="#accordion" href="#7"> | ||
| + | <div class="panel-title"> | ||
| + | <h3 style="padding: 0px; margin: 0px;">Parts construction</h3> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-6"> | ||
| + | Parts: | ||
| + | <ul style="list-style: none; padding: 0px; margin: 0px;"> | ||
| + | <li><strong>26-Cd</strong> (BBa_K1773022)</li> | ||
| + | <li><strong>27-Cd</strong> (BBa_K1773023)</li> | ||
| + | <li><strong>28-Cd</strong> (BBa_K1773024)</li> | ||
| + | </ul> | ||
| + | </div> | ||
| + | <div class="col-md-6"> | ||
| + | Written by: <strong>IR</strong><br /> | ||
| + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>BB</strong>, <strong>IS</strong> | ||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | </a> | ||
| + | </div> | ||
| + | <div id="7" class="panel-collapse collapse"> | ||
| + | <div class="panel-body"> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>About</strong></h3> | ||
| + | <p> | ||
| + | 26-Cd - Cascade with SRBS<br /> | ||
| + | 27-Cd - Cascade with MRBS<br /> | ||
| + | 28-Cd - Cascade with WRBS<br /> | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <table class="table"> | ||
| + | <thead> | ||
| + | <tr> | ||
| + | <th>Date</th> | ||
| + | <th>Materials</th> | ||
| + | <th>Experiment</th> | ||
| + | </tr> | ||
| + | </thead> | ||
| + | <tbody> | ||
| + | <tr> | ||
| + | <td>08 15</td> | ||
| + | <td>Cd1</td> | ||
| + | <td>Mutated Cd with Preffix and Suffix restriction with EcoRI and XbaI. L35 is produced.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 15</td> | ||
| + | <td> | ||
| + | P26<br /> | ||
| + | P27<br /> | ||
| + | P28<br /> | ||
| + | </td> | ||
| + | <td>PCR products from oligos restriction with EcoRI and SpeI. L26, L27, L28 and L29 are produced.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 15</td> | ||
| + | <td> | ||
| + | L35<br /> | ||
| + | L26<br /> | ||
| + | L27<br /> | ||
| + | L28<br /> | ||
| + | </td> | ||
| + | <td> | ||
| + | Ligation of: L35 and L26, L35 and L27, L35 and L28, <br />and their transformation into DH5α. | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 17 - <br /> - 08 20</td> | ||
| + | <td></td> | ||
| + | <td> | ||
| + | L35+L26 colony PCR with ST15 and VR.<br /> | ||
| + | L35+L27 colony PCR with ST15 and VR<br /> | ||
| + | L35+L28 colony PCR with ST15 and VR<br /> | ||
| + | 1<sup>st</sup> and 2<sup>nd</sup> colonies are inoculated.<br /> | ||
| + | 26-Cd, 27-Cd and 28-Cd are produced.<br /> | ||
| + | 26-Cd, 27-Cd and 28-Cd restriction with NspI.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 21</td> | ||
| + | <td></td> | ||
| + | <td>26-Cd, 27-Cd and 28-Cd sequencing.</td> | ||
| + | </tr> | ||
| + | </tbody> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>Conclusions and Results:</strong></h3> | ||
| + | <p style="height: auto; margin: auto;">07 31 – all of the colonies are successful.<br /> | ||
| + | 08 10 – all biobricks are sequenced succesfully.</p> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | <!-- /MethodBox --> | ||
| + | |||
| + | <!-- MethodBox --> | ||
| + | <div class="panel panel-default" style="position: relative"> | ||
| + | <div class="panel-heading"> | ||
| + | <a class="accordion-toggle collapsed" data-toggle="collapse" data-parent="#accordion" href="#8"> | ||
| + | <div class="panel-title"> | ||
| + | <h3 style="padding: 0px; margin: 0px;">Parts construction</h3> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-6"> | ||
| + | Part: <strong>P48</strong> (BBa_K1773026) | ||
| + | </div> | ||
| + | <div class="col-md-6"> | ||
| + | Written by: <strong>IR</strong><br /> | ||
| + | Performed by: <strong>ŠT</strong>, <strong>IR</strong>, <strong>IS</strong>, <strong>BB</strong> | ||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | </a> | ||
| + | </div> | ||
| + | <div id="8" class="panel-collapse collapse"> | ||
| + | <div class="panel-body"> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>About</strong></h3> | ||
| + | <p>pLac-cI repressor+ pLux/Ci-LuxI</p> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <table class="table"> | ||
| + | <thead> | ||
| + | <tr> | ||
| + | <th>Date</th> | ||
| + | <th>Materials</th> | ||
| + | <th>Experiment</th> | ||
| + | </tr> | ||
| + | </thead> | ||
| + | <tbody> | ||
| + | <tr> | ||
| + | <td>07 19</td> | ||
| + | <td>pLac (R0010)<br /> cI lambda repressor (P0151)</td> | ||
| + | <td>pLac biobrick was cleaved with EcoRI and SpeI, L10 was constructed.<br /> | ||
| + | cI Lambda repressor biobrick was digested with EcoRI and XbaI + FastAP, L11 was constructed.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>07 19</td> | ||
| + | <td>L10<br />L11</td> | ||
| + | <td>L11 and L10 were ligated and transformed to DH5α.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>07 20</td> | ||
| + | <td></td> | ||
| + | <td> | ||
| + | Colony PCR was performed with VF2 and VR primers.<br /> | ||
| + | 1<sup>st</sup> and 2<sup>nd</sup> colonies are inoculated. P18 is produced.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>07 21</td> | ||
| + | <td>P14.2<br />P18</td> | ||
| + | <td>P14.2 is digested with EcoRI and XbaI + Fast AP.<br /> | ||
| + | P18 is digested with EcoRI + SpeI.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>07 21</td> | ||
| + | <td></td> | ||
| + | <td>Restriction products are ligated and transformed into DH5α competent cells.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>07 22</td> | ||
| + | <td></td> | ||
| + | <td> | ||
| + | Colony PCR was performed with VF2 and VR primers.<br /> | ||
| + | 1<sup>st</sup> and 2<sup>nd</sup> colonies are inoculated. P48 is produced and sequenced. | ||
| + | </td> | ||
| + | </tr> | ||
| + | </tbody> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>Conclusions and Results:</strong></h3> | ||
| + | <p style="height: auto; margin: auto;"> 07 25 - P48 is sequenced successfully.</p> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | <!-- /MethodBox --> | ||
| + | |||
| + | <!-- MethodBox --> | ||
| + | <div class="panel panel-default" style="position: relative"> | ||
| + | <div class="panel-heading"> | ||
| + | <a class="accordion-toggle collapsed" data-toggle="collapse" data-parent="#accordion" href="#9"> | ||
| + | <div class="panel-title"> | ||
| + | <h3 style="padding: 0px; margin: 0px;">System characterization</h3> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-6"> | ||
| + | Parts: | ||
| + | <ul style="list-style: none; padding: 0px; margin: 0px;"> | ||
| + | <li><strong>Cd21L26</strong> (BBa_K1773022)</li> | ||
| + | <li><strong>Cd21L27</strong> (BBa_K1773023)</li> | ||
| + | <li><strong>Cd21L28</strong> (BBa_K1773024)</li> | ||
| + | </ul> | ||
| + | </div> | ||
| + | <div class="col-md-6"> | ||
| + | Written by: <strong>BB</strong><br /> | ||
| + | Performed by: <strong>BB</strong> | ||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | </a> | ||
| + | </div> | ||
| + | <div id="9" class="panel-collapse collapse"> | ||
| + | <div class="panel-body"> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>About</strong></h3> | ||
| + | <p> | ||
| + | Cd21L26 - Cascade with pLux/cI right promoter and strong RBS <br /> | ||
| + | Cd21L27 - Cascade with pLux/cI right promoter and Medium RBS <br /> | ||
| + | Cd21L28 - Cascade with pLux/cI right promoter and weak RBS <br /> | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <table class="table"> | ||
| + | <thead> | ||
| + | <tr> | ||
| + | <th>Date</th> | ||
| + | <th>Materials</th> | ||
| + | <th>Experiment</th> | ||
| + | </tr> | ||
| + | </thead> | ||
| + | <tbody> | ||
| + | <tr> | ||
| + | <td>08 26</td> | ||
| + | <td> | ||
| + | BL21-DE3<br /> | ||
| + | Cd21L26 <br /> | ||
| + | Cd21L27<br /> | ||
| + | Cd21L28<br /> | ||
| + | PET-HCsy3</td> | ||
| + | <td> | ||
| + | Transformation of Cd21L26 and PET-HCsy3 plasmids into BL21-DE3 bacteria. <br /> | ||
| + | Transformation of Cd21L27 and PET-HCsy3 plasmids into BL21-DE3 bacteria. <br /> | ||
| + | Transformation of Cd21L28 and PET-HCsy3 plasmids into BL21-DE3 bacteria. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 27</td> | ||
| + | <td> | ||
| + | BL21-DE3<br /> | ||
| + | Cd21L26 <br /> | ||
| + | Cd21L27<br /> | ||
| + | Cd21L28<br /> | ||
| + | PET-HCsy3</td> | ||
| + | <td> | ||
| + | Transformation of Cd21L26 plasmid into BL21-DE3 bacteria. <br /> | ||
| + | Transformation of Cd21L27 plasmid into BL21-DE3 bacteria. <br /> | ||
| + | Transformation of Cd21L28 plasmid into BL21-DE3 bacteria. <br /> | ||
| + | Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 28</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants <br /> | ||
| + | Cd21L26 <br /> | ||
| + | Cd21L27 <br /> | ||
| + | Cd21L28 <br /> | ||
| + | PET-HCsy3 </td> | ||
| + | <td> | ||
| + | Transformation of Cd21L26 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). <br /> | ||
| + | Transformation of Cd21L27 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). <br /> | ||
| + | Transformation of Cd21L28 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). <br /> | ||
| + | Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L26). <br /> | ||
| + | Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L27). <br /> | ||
| + | Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L28). <br /> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 29</td> | ||
| + | <td>BL21-DE3 transformants</td> | ||
| + | <td> | ||
| + | Growth of transformants in liquid LB. <br /> | ||
| + | Preparing to repeat transformation.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 30</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants <br /> | ||
| + | Cd21L26 <br /> | ||
| + | Cd21L27 <br /> | ||
| + | Cd21L28<br /> | ||
| + | PET-HCsy3</td> | ||
| + | <td> | ||
| + | Transformation of Cd21L26 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). <br /> | ||
| + | Transformation of Cd21L27 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). <br /> | ||
| + | Transformation of Cd21L28 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). <br /> | ||
| + | Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L26). <br /> | ||
| + | Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L27). <br /> | ||
| + | Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L28).</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 31</td> | ||
| + | <td>BL21-DE3 transformants</td> | ||
| + | <td> | ||
| + | Repeating transformation for better results.<br /> | ||
| + | Inoculating night culture of transformants.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 01</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants <br /> | ||
| + | Cd21L26 <br /> | ||
| + | Cd21L27 <br /> | ||
| + | Cd21L28 <br /> | ||
| + | PET-HCsy3</td> | ||
| + | <td> | ||
| + | Repeating transformation of Cd21L26 and PET-HCsy3 plasmids into BL21-DE3 bacteria, Cd21L27 and PET-HCsy3 and Cd21L28 and PET-HCsy3 into BL21-DE3 bacteria.<br /> | ||
| + | Repeating transformation of Cd21L26, Cd21L27 and Cd21L28 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). <br /> | ||
| + | Repeating transformation of PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L26, Cd21L27 and Cd21L28). <br /> | ||
| + | Trying to improve transformation by putting less antibiotics on a Petri dish.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 02</td> | ||
| + | <td>BL21-DE3 transformants</td> | ||
| + | <td> | ||
| + | Checking viability of the colonies by inoculating them in liquid LB for a few hours.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 03</td> | ||
| + | <td>BL21-DE3 transformants</td> | ||
| + | <td> | ||
| + | Transformants are inoculated into 20 mL LB and grown for 3 hours at 37ºC. <br /> | ||
| + | Half of the transformants are put into 16ºC and grown overnight. <br /> | ||
| + | Other half is centrifuged, supernatant discarded and cells put into 4ºC for further experiments.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 04</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants <br /> | ||
| + | Depleting buffer <br /> | ||
| + | PMSF <br /> | ||
| + | Protein Dye</td> | ||
| + | <td> | ||
| + | Transformants from 16ºC are centrifuged, supernatant discarded.<br /> | ||
| + | Lysing all transformant cells with ultrasound. <br /> | ||
| + | Lysate transferred into fresh collection tubes (1.5 mL) and used for Western Blot samples preparation. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 07</td> | ||
| + | <td> | ||
| + | Polyacrylamid gel <br /> | ||
| + | Protein samples</td> | ||
| + | <td> | ||
| + | SDS-PAGE is made in order to decide the right volume of protein solution for Western Blot.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 08</td> | ||
| + | <td> | ||
| + | Polyacrylamid gel <br /> | ||
| + | PVDF membraine <br /> | ||
| + | Primary antibodies</td> | ||
| + | <td> | ||
| + | Doing Western Blot <br /> | ||
| + | Incubating with primary antibodies overnight</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 10</td> | ||
| + | <td>Secondary antibodies</td> | ||
| + | <td> | ||
| + | Finishing Western Blot <br /> | ||
| + | Incubating with secondary antibodies <br /> | ||
| + | Visualizing blot results</td> | ||
| + | </tr> | ||
| + | </tbody> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>Conclusions and Results:</strong></h3> | ||
| + | <p style="height: auto; margin: auto;"> | ||
| + | 08 27 – Transformation unsuccessful. <br /> | ||
| + | 08 28 – Transformation successful. <br /> | ||
| + | 08 29 – Few colonies grown, but none multiplied in liquid LB with antibiotics. <br /> | ||
| + | 08 31 – Few colonies have grown on a plate. <br /> | ||
| + | 09 01 – Night culture did not grow. Transformation unsuccessful. <br /> | ||
| + | 09 02 – Few colonies have grown, but all viable. <br /> | ||
| + | 09 04 – Western Blot samples were made. <br /> | ||
| + | 09 07 – Decided what volume of protein should be used for western blot. <br /> | ||
| + | 09 10 – Visualized Western Blot results.</p> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | </html> | ||
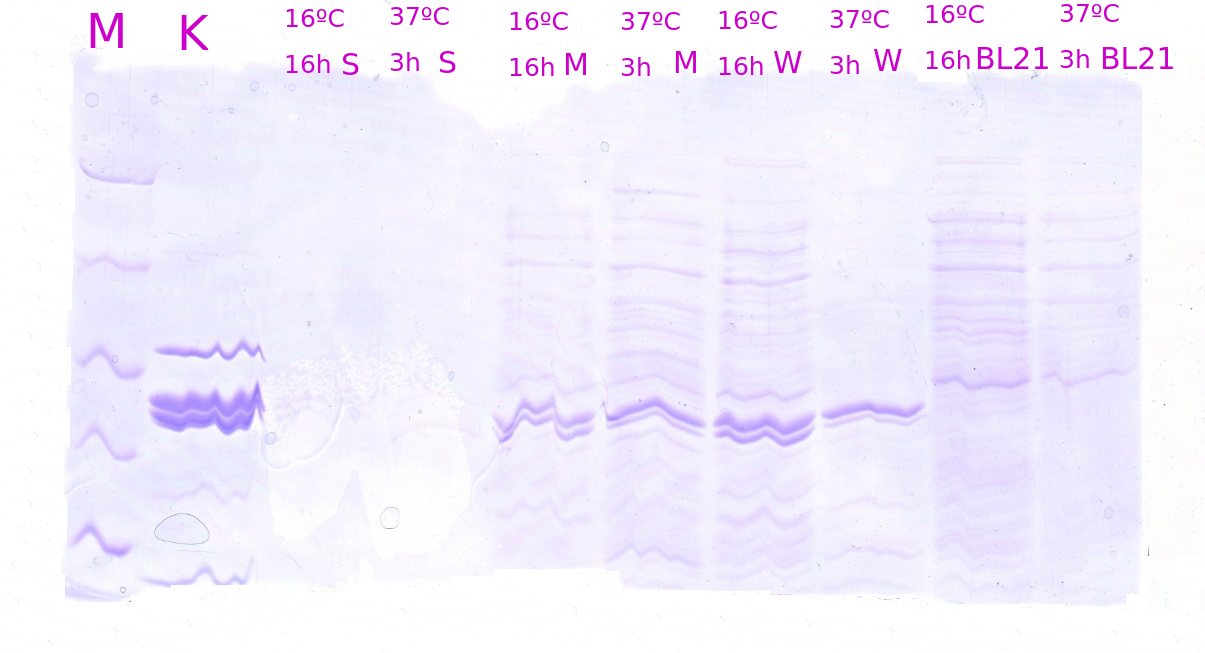
| + | [[File:Vilnius15_Igem_NDS_Page087_1.png|500px|thumb|<strong>Figure 1.</strong>SDS-PAGE of the Cascade biobricks. Expression of different Cascade biobricks were analysed in two incubation conditions after reaching a cell OD500 ~0.6. Biobricks incubated at 16 °C for 16hours showed a bit stronger expression strength rather than incubated at 37°C for 3hours. M-molecular mass marker (Spectra broad range protein protein ladder); K - possitive control of Csy3 protein; Cascade biobricks were expressed with strong (S), medium (M) or weak (W) RBS sites. BL - BL21-DE3 strain with no transformed plasmids.|center]] | ||
| + | <html> | ||
| + | |||
| + | </html> | ||
| + | [[File:Vilnius15_blotas_su_paaiskinimu.png|500px|thumb|<strong>Figure 2.</strong>Western blot of the Cascade biobricks. Expression of different Cascade biobricks were analysed in two incubation conditions after reaching a cell OD500 ~0.6. Biobricks incubated at 16 °C for 16hours showed a bit stronger expression strength rather than incubated at 37°C for 3hours. M-molecular mass marker (Spectra broad range protein protein ladder); K - possitive control of Csy3 protein; Cascade biobricks were expressed with strong (S), medium (M) or weak (W) RBS sites. BL - BL21-DE3 strain with no transformed plasmids|center]] | ||
| + | <html> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | <!-- /MethodBox --> | ||
| + | |||
| + | <!-- MethodBox --> | ||
| + | <div class="panel panel-default" style="position: relative"> | ||
| + | <div class="panel-heading"> | ||
| + | <a class="accordion-toggle collapsed" data-toggle="collapse" data-parent="#accordion" href="#10"> | ||
| + | <div class="panel-title"> | ||
| + | <h3 style="padding: 0px; margin: 0px;">System characterization</h3> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-6"> | ||
| + | Parts: | ||
| + | <ul style="list-style: none; padding: 0px; margin: 0px;"> | ||
| + | <li><strong>Cd21L26</strong> (BBa_K1773022)</li> | ||
| + | <li><strong>Cd21L27</strong> (BBa_K1773023)</li> | ||
| + | <li><strong>Cd21L28</strong> (BBa_K1773024)</li> | ||
| + | </ul> | ||
| + | </div> | ||
| + | <div class="col-md-6"> | ||
| + | Written by: <strong>BB</strong><br /> | ||
| + | Performed by: <strong>BB</strong> and <strong>IO</strong> | ||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | </a> | ||
| + | </div> | ||
| + | <div id="10" class="panel-collapse collapse"> | ||
| + | <div class="panel-body"> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>About</strong></h3> | ||
| + | <p> | ||
| + | Cd21L26 - Cascade with pLux/cI right promoter and strong RBS <br /> | ||
| + | Cd21L27 - Cascade with pLux/cI right promoter and Medium RBS <br /> | ||
| + | Cd21L28 - Cascade with pLux/cI right promoter and weak RBS <br /> | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <table class="table"> | ||
| + | <thead> | ||
| + | <tr> | ||
| + | <th>Date</th> | ||
| + | <th>Materials</th> | ||
| + | <th>Experiment</th> | ||
| + | </tr> | ||
| + | </thead> | ||
| + | <tbody> | ||
| + | <tr> | ||
| + | <td>09 04</td> | ||
| + | <td> | ||
| + | SP1<br /> | ||
| + | SP3<br /> | ||
| + | pAcYc plasmid<br /> | ||
| + | DH5α<br /> | ||
| + | Karbenicilin</td> | ||
| + | <td> | ||
| + | Restriction of pCDF vector, SP1 and SP3 genes with EcoRI and XbaI. <br /> | ||
| + | Ligating pieces afterwards, transforming new pCDF plasmid into bacteria. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 05</td> | ||
| + | <td> | ||
| + | DH5α transformants<br /> | ||
| + | Bl21-DE3 transformants<br /> | ||
| + | pCola-Cas1-Cas3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Karbenicilin </td> | ||
| + | <td> | ||
| + | Colony PCR. Inoculating good colonies.<br /> | ||
| + | Inoculating Bl21-DE3 transformants (BL21-DE3 with Cd21L26 + HCsy3; Cd21L27 + HCsy3; Cd21L28 + HCsy3) overnight. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 06</td> | ||
| + | <td> | ||
| + | Bl21-DE3 transformants<br /> | ||
| + | pCola-Cas1-Cas3 plasmid<br /> | ||
| + | SP1<br /> | ||
| + | SP3<br /> | ||
| + | pCDF<br /> | ||
| + | DH5α bacteria<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin<br /> | ||
| + | Streptomycin </td> | ||
| + | <td> | ||
| + | Inoculating night culture of Bl21-DE3 transformants for 3 hours. <br /> | ||
| + | Preparing competent cells. Transforming pCola plasmid into BL21-DE3 transformants. <br /> | ||
| + | Repeat cloning of SP1 and SP3 into pCDF vector and transforming them into DH5α. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 07</td> | ||
| + | <td> | ||
| + | DH5α transformants<br /> | ||
| + | Bl21-DE3 transformants<br /> | ||
| + | pCola-Cas1-Cas3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Karbenicilin <br /> | ||
| + | Kanamycin <br /> | ||
| + | Streptomycin </td> | ||
| + | <td> | ||
| + | Colony PCR. Inoculating good SP1 and SP3 colonies overnight. | ||
| + | Repeating yesterdays transformation. Inoculating night colonies again, in case of transformation failure. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 08</td> | ||
| + | <td> | ||
| + | DH5α transformants<br /> | ||
| + | Bl21-DE3 transformants<br /> | ||
| + | pCola-Cas1-Cas3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Karbenicilin <br /> | ||
| + | Kanamycin </td> | ||
| + | <td> | ||
| + | Extracting SP1 and SP3 plasmids. Repeating transformation. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 09</td> | ||
| + | <td> | ||
| + | Bl21-DE3 bacteria<br /> | ||
| + | pCola-Cas1-Cas3 plasmid<br /> | ||
| + | Cd21L26 plasmid<br /> | ||
| + | Cd21L27 plasmid<br /> | ||
| + | Cd21L28 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin </td> | ||
| + | <td> | ||
| + | Transformation of pCola and Cd21L26 into BL21-DE3 bacteria.<br /> | ||
| + | Transformation of pCola and Cd21L27 into BL21-DE3 bacteria.<br /> | ||
| + | Transformation of pCola and Cd21L28 into BL21-DE3 bacteria.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 10</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin </td> | ||
| + | <td>Inoculating transformants overnight. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 11</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants<br /> | ||
| + | pCDF-SP1 plasmid<br /> | ||
| + | pCDF-SP3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Measuring night culture optical dense. Equalizing optical dense of all bacteria.<br /> | ||
| + | Preparing cells for transformation. Transforming pCDF-SP1, pCDF-SP3 and pCDF plasmids into BL21-DE3 transformants. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 12</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants<br /> | ||
| + | pCDF-SP1 plasmid<br /> | ||
| + | pCDF-SP3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Repeating transformation. Before inoculating bacteria on a Petri dish, it is diluted 100 and 10000 times.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 13</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants<br /> | ||
| + | pCDF-SP1 plasmid<br /> | ||
| + | pCDF-SP3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Repeating transformation.<br /> | ||
| + | Before inoculating bacteria on a petri dish, it is diluted 100 and 1000 times. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 14</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants<br /> | ||
| + | pCDF-SP1 plasmid<br /> | ||
| + | pCDF-SP3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Repeating transformation. <br /> | ||
| + | Before bacteria inoculation on a petri dish, it is diluted 100 and 10000 times.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 15</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin</td> | ||
| + | <td> | ||
| + | Inoculating new BL21-DE3 transformant cells. Growing overnight. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 16</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants<br /> | ||
| + | pCDF-SP1 plasmid<br /> | ||
| + | pCDF-SP3 plasmid<br /> | ||
| + | Chloramphenicol<br /> | ||
| + | Kanamycin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Repeating transformation. Before bacteria inoculation on a petri dish, it is diluted 100 and 10000 times.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 17</td> | ||
| + | <td> | ||
| + | BL21-DE3 transformants </td> | ||
| + | <td> | ||
| + | Counting grown colonies. </td> | ||
| + | </tr> | ||
| + | </tbody> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>Conclusions and Results:</strong></h3> | ||
| + | <p style="height: auto; margin: auto;"> | ||
| + | 09 05 – Transformation successful. Colony PCR showed, that SP1 and SP3 fragments were cloned successfully. <br /> | ||
| + | 09 06 – Mistake found. SP1 and SP3 were cloned into a wrong vector. <br /> | ||
| + | 09 07 – Transformation was unsuccessful.<br /> | ||
| + | 09 08 – Transformation unsuccessful. Trying to reduce antibiotic quantity. <br /> | ||
| + | 09 09 – Transformation unsuccessful. PET-HCsy3 plasmid may have inhibited transformation. <br /> | ||
| + | 09 10 – Transformation successful. <br /> | ||
| + | 09 11 – BL21-DE3 transformed with Cd21L27+pCola did not grow. <br /> | ||
| + | 09 12 – Too much bacteria grew up. <br /> | ||
| + | 09 13 – Transformation unsuccessful. In some dishes still too many bacteria grown. <br /> | ||
| + | 09 14 – Transformation unsuccessful. <br /> | ||
| + | 09 15 – Transformation unsuccessful. Probably not enough pCDF-SP1 and pCDF-SP3 vector. <br /> | ||
| + | 09 17 - Transformation successful.</p> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | </html> | ||
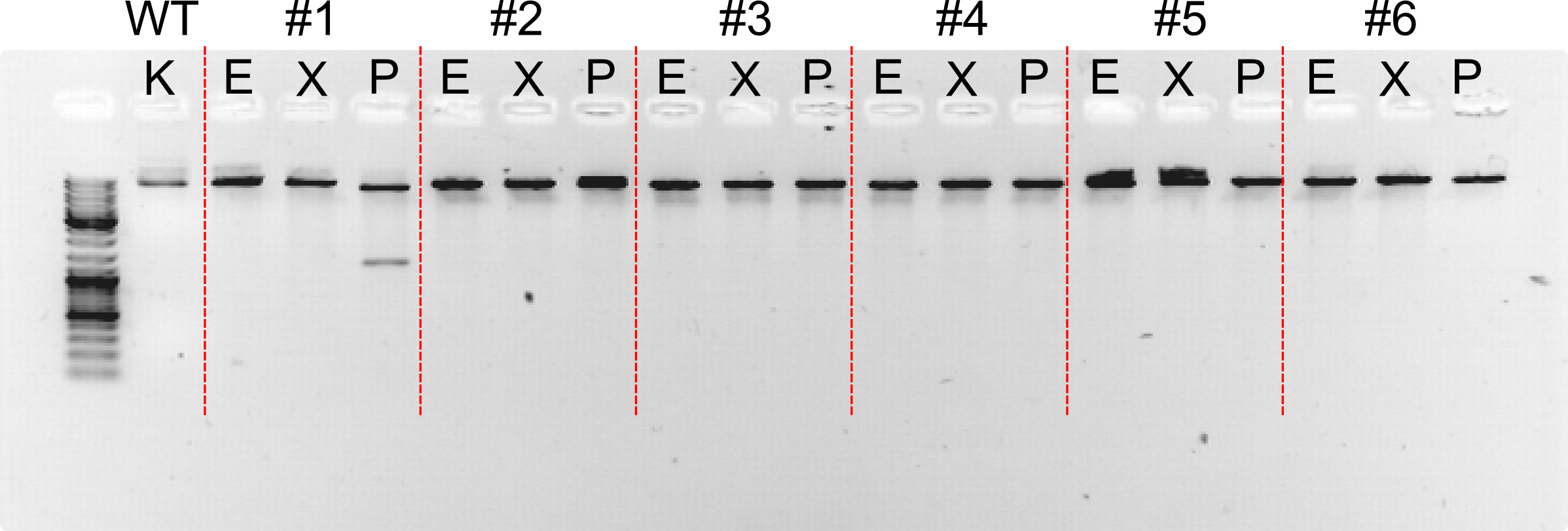
| + | [[File:Vilnius15_09_07_Sp1_Sp3_koloniju_PGR.png|500px|thumb|<strong>Figure 1</strong> SP1 and SP3 colony PCR. SP1 and SP3 genes were cloned into pCDF vector. After transformation colonies were checked with PCR. 7 colonie (well no. 7) of SP1 cloning and 3 colonie (well no. 12) of SP3 cloning were good. We can see a 700 bp fragment, that indicates the needed fragment. |center]] | ||
| + | <html> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | <!-- /MethodBox --> | ||
| + | |||
| + | <!-- MethodBox --> | ||
| + | <div class="panel panel-default" style="position: relative"> | ||
| + | <div class="panel-heading"> | ||
| + | <a class="accordion-toggle collapsed" data-toggle="collapse" data-parent="#accordion" href="#11"> | ||
| + | <div class="panel-title"> | ||
| + | <h3 style="padding: 0px; margin: 0px;">System characterization</h3> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-6"> | ||
| + | Part: <strong>P30</strong>(BBa_K1773007) | ||
| + | </div> | ||
| + | <div class="col-md-6"> | ||
| + | Written by: <strong>BB</strong><br /> | ||
| + | Performed by: <strong>BB</strong>, <strong>IS</strong>, <strong>IR</strong> and <strong>IO</strong> | ||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | </a> | ||
| + | </div> | ||
| + | <div id="11" class="panel-collapse collapse"> | ||
| + | <div class="panel-body"> | ||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>About</strong></h3> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <table class="table"> | ||
| + | <thead> | ||
| + | <tr> | ||
| + | <th>Date</th> | ||
| + | <th>Materials</th> | ||
| + | <th>Experiment</th> | ||
| + | </tr> | ||
| + | </thead> | ||
| + | <tbody> | ||
| + | <tr> | ||
| + | <td>08 26</td> | ||
| + | <td> | ||
| + | P20<br /> | ||
| + | Carbicilin</td> | ||
| + | <td> | ||
| + | Transformation of P20 into JM109. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 27</td> | ||
| + | <td> | ||
| + | P30<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Transformation into JM109 (transformed with P20) </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 28</td> | ||
| + | <td> | ||
| + | JM109 transformants<br /> | ||
| + | P30<br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Growing of JM109 transformants (low copy) and preparing for new transformation. <br /> | ||
| + | Transformation of P30 into empty JM109. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 29</td> | ||
| + | <td> | ||
| + | JM109 transformants<br /> | ||
| + | P30<br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Transformation of P30 into JM109 (transformed with P20) </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 30</td> | ||
| + | <td> | ||
| + | JM109 transformants<br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Transformation of P20 into JM109 (transformed with P30)</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>08 31</td> | ||
| + | <td> | ||
| + | JM109 transformants<br /> | ||
| + | P30<br /> | ||
| + | P20 <br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Transformation of P20 into JM109 (transformed with P30). <br /> | ||
| + | Transformation of P30 into JM109 (transformed with P20). </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 01</td> | ||
| + | <td> | ||
| + | JM109 transformants<br /> | ||
| + | JM109 (fresh)<br /> | ||
| + | P20<br /> | ||
| + | P30<br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Transformation of P20 and P30 into JM109 (fresh).<br /> | ||
| + | Transformation of P20 into JM109 (transformed with P30).<br /> | ||
| + | Transformation of P30 into JM109 (transformed with P20). </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 02</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)<br /> | ||
| + | IPTG<br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Growing transformants for 4 hours in a liquid LB. Transfer transformants into LB with different IPTG concentrations and grow overnight. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 12</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)</td> | ||
| + | <td>Fluorescence measurement and analysis. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 06</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)<br /> | ||
| + | IPTG<br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Growing transformants for 4 hours in a liquid LB. Transfer transformants into LB with IPTG and grow overnight.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 07</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)</td> | ||
| + | <td> | ||
| + | Fluorescence measurement and analysis.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 08</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)<br /> | ||
| + | IPTG<br /> | ||
| + | Carbicilin<br /> | ||
| + | Chloramphenicol</td> | ||
| + | <td> | ||
| + | Growing transformants for 4 hours in a liquid LB. Transfer transformants into LB with IPTG and grow overnight. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 10</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)</td> | ||
| + | <td> | ||
| + | Fluorescence measurement and analysis.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 11</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)</td> | ||
| + | <td> | ||
| + | Fluorescence measurement and analysis.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 13</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)<br /> | ||
| + | pCDF vector<br /> | ||
| + | Carbicilin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Growing JM109 transformants as competent cells.<br /> | ||
| + | Restriction and ligation of P30 fragment into pCDF vector. <br /> | ||
| + | Transformation of new P30 into JM109 (transformed with P20). </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 14</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)<br /> | ||
| + | Carbicilin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Colony PCR.</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 15</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)<br /> | ||
| + | Carbicilin<br /> | ||
| + | Streptomycin</td> | ||
| + | <td> | ||
| + | Inoculating bacteria for 4 hours. Inoculating overnight with IPTG. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 16</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)</td> | ||
| + | <td> | ||
| + | Measuring fluorescence. </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>09 17</td> | ||
| + | <td> | ||
| + | JM109 transformants (with P20 and P30)</td> | ||
| + | <td> | ||
| + | Measuring fluorescence. </td> | ||
| + | </tr> | ||
| + | </tbody> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | <div class="row"> | ||
| + | <div class="col-md-12"> | ||
| + | <h3><strong>Conclusions and Results:</strong></h3> | ||
| + | <p style="height: auto; margin: auto;"> | ||
| + | 08 26 – Transformation successful.<br /> | ||
| + | 08 27 – Transformation unsuccessful.<br /> | ||
| + | 08 29 – JM109 grown successfully. Transformation of P30 into empty JM109 was successful.<br /> | ||
| + | 08 30 – Transformation unsuccessful.<br /> | ||
| + | 08 31 - Transformation unsuccessful. The samples could have been pollen. <br /> | ||
| + | 09 01 – Transformation unsuccessful. Maybe it was too much antibiotic on a petri dish.<br /> | ||
| + | 09 02 – Transformation successful.<br /> | ||
| + | 09 03 – Bad fluorimeter options.<br /> | ||
| + | 09 07 – No repeated samples were made.<br /> | ||
| + | 09 11 – Fluorescence measurement seems successful.<br /> | ||
| + | 09 12 – Mistake found. P30 and P20 fragments are in an uncompetable plasmids. Repeating whole experiment. <br /> | ||
| + | 09 14 - Transformation successful. Colony PCR successful. <br /> | ||
| + | 09 17 - Fluorescence measurements are successful and conclusive.</td> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | </div> | ||
| + | </div> | ||
| + | </div> | ||
| + | <!-- /MethodBox --> | ||
</section> | </section> | ||
</body> | </body> | ||
| − | |||
</div> | </div> | ||
</div> | </div> | ||
| + | </html> | ||
{{Team:Vilnius-Lithuania/Vilnius15_Footer}} | {{Team:Vilnius-Lithuania/Vilnius15_Footer}} | ||
Latest revision as of 03:56, 19 September 2015
About
pLac+cI.
P10 and P11 plasmids are used. pLac (P11) and repressor cI (P10) ligation.
| Date | Materials | Experiment |
|---|---|---|
| 06 30 | P10 P11 |
P10 and P11 transformation into DH5α. |
| 07 01 | Bacteria inoculation (4 minipreps). | |
| 07 09 | P10 P11 |
Plasmids’ restriction (3 ug each). P10 with EcoRI and XBal (+FastAp). P11 with EcoRI and SpeI. Extraction from gel L10 and L11 are produced. Conc.: L11 – 2,4 ng/μL. |
| 07 13 | P10 |
Repeated restriction of P10 with EcoRI and XBal (+FastAp). Conc.: L10 – 45,3 ng/μL. L10 and L11 ligation. P14 is produced. |
| 07 14 | Bacteria with P14 | Colony PCR of P14 bacteria. 1st and 2nd colonies are inoculated. |
| P14.2 | Sequencing |
Conclusions and Results:
07 14 – all of the colonies are successful, except third colony.
About
Cas3 with mutated restriction sites.
L7+L15. Concentration - 261 ng/μL
| Date | Materials | Experiment |
|---|---|---|
| 06 30 | P13 P7 |
P13 and P7 transformation into DH5α. |
| 07 01 | Transformants are inoculated (4 minipreps each). | |
| 07 02 | Plasmid extraction. Concentrations: P7 – 174,1 ng/μL and 166 ng/μL. P13 - 216 ng/μL and 131, 5 ng/μL. |
|
| 07 13 | P13 | Cas3 mutagenesis (with ST1/2, ST3/4, ST5/6 primers). |
| 07 16 | Mutated P13 | Cas3 plasmids extraction. |
| 07 21-23 | Mutated P13 | Restriction with EcoRI, XbaI and PstI. |
| 07 27 | Mutated P13 | Cas3 mutagenesis. |
| 07 29 | P7 | Restriction with EcoRI and SpeI. L7 is produced. |
| 07 31 | P13 | |
| 08 02 | Mutated P13 | Mutated Cas3 restriction with EcoRI, XbaI and PstI. |
| 08 05 | Mutated P13 | Cas3 mutagenesis. |
| 08 07 | Mutated P13 | Mutated Cas3 restriction with PstI. Mutagenesis was successful. L15 is produced. |
| 08 11 | L7 L15 |
L7 and L15 ligation with pSB1C3 (EcoRI+XBal+FastAp). P15.1 is produced. |
| P15.1 | Sequencing. |
Conclusions and Results:
07 21 – Mutagenesis was unsuccessful.
08 02 – successfully mutagenized EcoRI and XbaI sites.
08 07 – successful mutagenesis confirmed (XbaI and EcoRI sites are mutated).
08 07 – Successful mutagenesis (PstI, XbaI and EcoRI sites are mutated).
About
P26 – L7+L26 pLux/cI+SRBS
P27 – L7+L27 pLux/cI+MRBS
P28 – L7+L28 pLux/cI+WRBS
P29 – L7+L29 pLux/cI
| Date | Materials | Experiment |
|---|---|---|
| 07 27 |
ST9 ST10 ST11 ST12 ST15 ST16 ST17 ST18 ST19 ST20 |
pLux/cI contruction from oligos – PCR reaction with ST9 and ST10/11/12 primers, then with ST15 and ST16/17/18. Parallel PCR with ST9 and ST19 and then with ST15 and ST20. PCR products purification. |
| 07 28 | PCR products from oligos restriction with EcoRI and SpeI. L26, L27, L28 and L29 are produced. |
|
| 07 28 |
L7 L26 L27 L28 L29 |
Ligation of: L7 and L26, L7 and L27, L7 and L28, L7 and L29 and their transformation into DH5α. |
| 07 29 - - 08 05 |
L7+L26 colony PCR with ST15 and VR. L7+L27 colony PCR with ST15 and VR. L7+L28 colony PCR with ST15 and VR. 1st and 2nd colonies are inoculated. P26, P27, 28 and P29 are produced P26, P27, 28 and P29 restriction with NspI. |
|
| 08 10 | P26, P27, 28 and P29 sequencing with VF2. |
Conclusions and Results:
07 31 – all of the colonies are successful.
08 10 – all biobricks are sequenced succesfully.
About
cI repressor promoter (BBa_R1051) (P4) + screen plasmid intermediate (GFP gene) (BBa_I13504) (P5).
| Date | Materials | Experiment |
|---|---|---|
| 06 30 | P14 | P4 transformation into DH5α. |
| 07 01 | Bacteria inoculation (4 minipreps). | |
| 07 02 | P4 |
Plasmid extraction. Concentrations: P4 - 238,3 ng/μL and 224,3 ng/μL. P4 restriction with EcoRI and XBal. Extraction from gel. L4 is produced (con. 16,2 ng/μL). |
| 07 07 | P5 |
P5 transformation into DH5α. Bacteria inoculation. |
| 07 08 | P5 plasmid extraction (P5.1, P5.2, P5.3, P5.4, P5.5 are produced). | |
| 07 29 | P5.1 and P5.3 |
Restriction with EcoRI and SpeI. Extraction from gel. L5.1A, L5.1B, L5.1C and L5.3A, L5.3B, L5.3C are produced. Conc.: L5.1A – 0,6 ng/μL, L5.1B – 0,6 ng/μL, L5.1C – 2,4 ng/μL, L5.3A – 0,7 ng/μL, L5.3B - 3 ng/μL, L5.3C – 5,1 ng/μL. Ligation of L4 and L5.1, L4 and L5.3. P30 is produced. |
| 08 01 | P30 | Sequencing. |
Conclusions and Results:
07 09 – fragments were not the right ones.
07 29 – the right fragments are produced.
About
P35 - pLux/cI+SRBS(Strong RBS)+Cas3 (L26+L15.1)
P36 - pLux/cI+MRBS(Medium RBS)+Cas3 (L27+L15.1)
P37 - pLux/cI+WRBS(Weak RBS)+Cas3 (L28+L15.1)
P26 – L7+L26 (pLux/cI+SRBS)
P27 - pLux/cI+MRBS(Medium RBS)
P28 - pLux/cI+WRBS(Weak RBS)
| Date | Materials | Experiment |
|---|---|---|
| 07 28 | P15 | Mutated Cas3 with Preffix and Suffix restriction with EcoRI and XbaI. L15 is produced. |
| 07 28 |
P26 P27 P28 |
PCR products from oligos restriction with EcoRI and SpeI. L26, L27, L28 and L29 are produced. |
| 07 28 |
L15 L26 L27 L28 |
Ligation of: L15 and L26, L15 and L27, L15 and L28, and their transformation into DH5α. |
| 07 29 - - 08 05 |
L7+L26 colony PCR with ST15 and VR. L7+L27 colony PCR with ST15 and VR L7+L28 colony PCR with ST15 and VR 1st and 2nd colonies are inoculated. P35, P36 and P37 are produced P35, P36, P37 restriction with NspI. |
|
| 08 10 | P35, P36, P37 sequencing. |
Conclusions and Results:
07 31 – all of the colonies are successful.
08 10 – all biobricks are sequenced succesfully.
About
Mutated Cascade protein.
| Date | Materials | Experiment |
|---|---|---|
| 06 30 | Cd | Cd transformation into DH5α. |
| 07 01 | Transformants are inoculated (4 minipreps each). | |
| 07 02 | Plasmid extraction. Conc.: 329 ng/μL and 294 ng/μL. | |
| 07 14 | Cd | Cd mutagenesis (with ST13/14 primers). |
| 07 16 - 17 | Mutated plasmids extraction. Restriction with EcoRI. | |
| 07 19 | Mutated Cd | Restriction repeat. |
| 07 21 | Mutated Cd | Repeated mutagenesis (with ST13/14). |
| 07 27 | Mutated Cd | Mutated plasmids extraction. Mutated Cd restriction. |
| 08 10 | Mutated Cd L7 |
PCR with mutated Cd with ST26/27. PCR product purification. Conc.: 63 ng/μL. Restriction with EcoRI and SpeI. Mutated Cd ligation with L7. Cd1 is produced. |
| Cd1 | Sequencing. |
Conclusions and Results:
07 16-17 – Mutagenesis was unsuccessful.
07 27 – Mutagenesis conformed as a successful.
About
26-Cd - Cascade with SRBS
27-Cd - Cascade with MRBS
28-Cd - Cascade with WRBS
| Date | Materials | Experiment |
|---|---|---|
| 08 15 | Cd1 | Mutated Cd with Preffix and Suffix restriction with EcoRI and XbaI. L35 is produced. |
| 08 15 |
P26 P27 P28 |
PCR products from oligos restriction with EcoRI and SpeI. L26, L27, L28 and L29 are produced. |
| 08 15 |
L35 L26 L27 L28 |
Ligation of: L35 and L26, L35 and L27, L35 and L28, and their transformation into DH5α. |
| 08 17 - - 08 20 |
L35+L26 colony PCR with ST15 and VR. L35+L27 colony PCR with ST15 and VR L35+L28 colony PCR with ST15 and VR 1st and 2nd colonies are inoculated. 26-Cd, 27-Cd and 28-Cd are produced. 26-Cd, 27-Cd and 28-Cd restriction with NspI. |
|
| 08 21 | 26-Cd, 27-Cd and 28-Cd sequencing. |
Conclusions and Results:
07 31 – all of the colonies are successful.
08 10 – all biobricks are sequenced succesfully.
About
pLac-cI repressor+ pLux/Ci-LuxI
| Date | Materials | Experiment |
|---|---|---|
| 07 19 | pLac (R0010) cI lambda repressor (P0151) |
pLac biobrick was cleaved with EcoRI and SpeI, L10 was constructed. cI Lambda repressor biobrick was digested with EcoRI and XbaI + FastAP, L11 was constructed. |
| 07 19 | L10 L11 |
L11 and L10 were ligated and transformed to DH5α. |
| 07 20 |
Colony PCR was performed with VF2 and VR primers. 1st and 2nd colonies are inoculated. P18 is produced. |
|
| 07 21 | P14.2 P18 |
P14.2 is digested with EcoRI and XbaI + Fast AP. P18 is digested with EcoRI + SpeI. |
| 07 21 | Restriction products are ligated and transformed into DH5α competent cells. | |
| 07 22 |
Colony PCR was performed with VF2 and VR primers. 1st and 2nd colonies are inoculated. P48 is produced and sequenced. |
Conclusions and Results:
07 25 - P48 is sequenced successfully.
About
Cd21L26 - Cascade with pLux/cI right promoter and strong RBS
Cd21L27 - Cascade with pLux/cI right promoter and Medium RBS
Cd21L28 - Cascade with pLux/cI right promoter and weak RBS
| Date | Materials | Experiment |
|---|---|---|
| 08 26 |
BL21-DE3 Cd21L26 Cd21L27 Cd21L28 PET-HCsy3 |
Transformation of Cd21L26 and PET-HCsy3 plasmids into BL21-DE3 bacteria. Transformation of Cd21L27 and PET-HCsy3 plasmids into BL21-DE3 bacteria. Transformation of Cd21L28 and PET-HCsy3 plasmids into BL21-DE3 bacteria. |
| 08 27 |
BL21-DE3 Cd21L26 Cd21L27 Cd21L28 PET-HCsy3 |
Transformation of Cd21L26 plasmid into BL21-DE3 bacteria. Transformation of Cd21L27 plasmid into BL21-DE3 bacteria. Transformation of Cd21L28 plasmid into BL21-DE3 bacteria. Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria. |
| 08 28 |
BL21-DE3 transformants Cd21L26 Cd21L27 Cd21L28 PET-HCsy3 |
Transformation of Cd21L26 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). Transformation of Cd21L27 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). Transformation of Cd21L28 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L26). Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L27). Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L28). |
| 08 29 | BL21-DE3 transformants |
Growth of transformants in liquid LB. Preparing to repeat transformation. |
| 08 30 |
BL21-DE3 transformants Cd21L26 Cd21L27 Cd21L28 PET-HCsy3 |
Transformation of Cd21L26 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). Transformation of Cd21L27 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). Transformation of Cd21L28 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L26). Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L27). Transformation of and PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L28). |
| 08 31 | BL21-DE3 transformants |
Repeating transformation for better results. Inoculating night culture of transformants. |
| 09 01 |
BL21-DE3 transformants Cd21L26 Cd21L27 Cd21L28 PET-HCsy3 |
Repeating transformation of Cd21L26 and PET-HCsy3 plasmids into BL21-DE3 bacteria, Cd21L27 and PET-HCsy3 and Cd21L28 and PET-HCsy3 into BL21-DE3 bacteria. Repeating transformation of Cd21L26, Cd21L27 and Cd21L28 plasmid into BL21-DE3 bacteria (transformed with PET-HCsy3). Repeating transformation of PET-HCsy3 plasmid into BL21-DE3 bacteria (transformed with Cd21L26, Cd21L27 and Cd21L28). Trying to improve transformation by putting less antibiotics on a Petri dish. |
| 09 02 | BL21-DE3 transformants | Checking viability of the colonies by inoculating them in liquid LB for a few hours. |
| 09 03 | BL21-DE3 transformants |
Transformants are inoculated into 20 mL LB and grown for 3 hours at 37ºC. Half of the transformants are put into 16ºC and grown overnight. Other half is centrifuged, supernatant discarded and cells put into 4ºC for further experiments. |
| 09 04 |
BL21-DE3 transformants Depleting buffer PMSF Protein Dye |
Transformants from 16ºC are centrifuged, supernatant discarded. Lysing all transformant cells with ultrasound. Lysate transferred into fresh collection tubes (1.5 mL) and used for Western Blot samples preparation. |
| 09 07 |
Polyacrylamid gel Protein samples |
SDS-PAGE is made in order to decide the right volume of protein solution for Western Blot. |
| 09 08 |
Polyacrylamid gel PVDF membraine Primary antibodies |
Doing Western Blot Incubating with primary antibodies overnight |
| 09 10 | Secondary antibodies |
Finishing Western Blot Incubating with secondary antibodies Visualizing blot results |
Conclusions and Results:
08 27 – Transformation unsuccessful.
08 28 – Transformation successful.
08 29 – Few colonies grown, but none multiplied in liquid LB with antibiotics.
08 31 – Few colonies have grown on a plate.
09 01 – Night culture did not grow. Transformation unsuccessful.
09 02 – Few colonies have grown, but all viable.
09 04 – Western Blot samples were made.
09 07 – Decided what volume of protein should be used for western blot.
09 10 – Visualized Western Blot results.

About
Cd21L26 - Cascade with pLux/cI right promoter and strong RBS
Cd21L27 - Cascade with pLux/cI right promoter and Medium RBS
Cd21L28 - Cascade with pLux/cI right promoter and weak RBS
| Date | Materials | Experiment |
|---|---|---|
| 09 04 |
SP1 SP3 pAcYc plasmid DH5α Karbenicilin |
Restriction of pCDF vector, SP1 and SP3 genes with EcoRI and XbaI. Ligating pieces afterwards, transforming new pCDF plasmid into bacteria. |
| 09 05 |
DH5α transformants Bl21-DE3 transformants pCola-Cas1-Cas3 plasmid Chloramphenicol Karbenicilin |
Colony PCR. Inoculating good colonies. Inoculating Bl21-DE3 transformants (BL21-DE3 with Cd21L26 + HCsy3; Cd21L27 + HCsy3; Cd21L28 + HCsy3) overnight. |
| 09 06 |
Bl21-DE3 transformants pCola-Cas1-Cas3 plasmid SP1 SP3 pCDF DH5α bacteria Chloramphenicol Kanamycin Streptomycin |
Inoculating night culture of Bl21-DE3 transformants for 3 hours. Preparing competent cells. Transforming pCola plasmid into BL21-DE3 transformants. Repeat cloning of SP1 and SP3 into pCDF vector and transforming them into DH5α. |
| 09 07 |
DH5α transformants Bl21-DE3 transformants pCola-Cas1-Cas3 plasmid Chloramphenicol Karbenicilin Kanamycin Streptomycin |
Colony PCR. Inoculating good SP1 and SP3 colonies overnight. Repeating yesterdays transformation. Inoculating night colonies again, in case of transformation failure. |
| 09 08 |
DH5α transformants Bl21-DE3 transformants pCola-Cas1-Cas3 plasmid Chloramphenicol Karbenicilin Kanamycin |
Extracting SP1 and SP3 plasmids. Repeating transformation. |
| 09 09 |
Bl21-DE3 bacteria pCola-Cas1-Cas3 plasmid Cd21L26 plasmid Cd21L27 plasmid Cd21L28 plasmid Chloramphenicol Kanamycin |
Transformation of pCola and Cd21L26 into BL21-DE3 bacteria. Transformation of pCola and Cd21L27 into BL21-DE3 bacteria. Transformation of pCola and Cd21L28 into BL21-DE3 bacteria. |
| 09 10 |
BL21-DE3 transformants Chloramphenicol Kanamycin |
Inoculating transformants overnight. |
| 09 11 |
BL21-DE3 transformants pCDF-SP1 plasmid pCDF-SP3 plasmid Chloramphenicol Kanamycin Streptomycin |
Measuring night culture optical dense. Equalizing optical dense of all bacteria. Preparing cells for transformation. Transforming pCDF-SP1, pCDF-SP3 and pCDF plasmids into BL21-DE3 transformants. |
| 09 12 |
BL21-DE3 transformants pCDF-SP1 plasmid pCDF-SP3 plasmid Chloramphenicol Kanamycin Streptomycin |
Repeating transformation. Before inoculating bacteria on a Petri dish, it is diluted 100 and 10000 times. |
| 09 13 |
BL21-DE3 transformants pCDF-SP1 plasmid pCDF-SP3 plasmid Chloramphenicol Kanamycin Streptomycin |
Repeating transformation. Before inoculating bacteria on a petri dish, it is diluted 100 and 1000 times. |
| 09 14 |
BL21-DE3 transformants pCDF-SP1 plasmid pCDF-SP3 plasmid Chloramphenicol Kanamycin Streptomycin |
Repeating transformation. Before bacteria inoculation on a petri dish, it is diluted 100 and 10000 times. |
| 09 15 |
BL21-DE3 transformants Chloramphenicol Kanamycin |
Inoculating new BL21-DE3 transformant cells. Growing overnight. |
| 09 16 |
BL21-DE3 transformants pCDF-SP1 plasmid pCDF-SP3 plasmid Chloramphenicol Kanamycin Streptomycin |
Repeating transformation. Before bacteria inoculation on a petri dish, it is diluted 100 and 10000 times. |
| 09 17 | BL21-DE3 transformants | Counting grown colonies. |
Conclusions and Results:
09 05 – Transformation successful. Colony PCR showed, that SP1 and SP3 fragments were cloned successfully.
09 06 – Mistake found. SP1 and SP3 were cloned into a wrong vector.
09 07 – Transformation was unsuccessful.
09 08 – Transformation unsuccessful. Trying to reduce antibiotic quantity.
09 09 – Transformation unsuccessful. PET-HCsy3 plasmid may have inhibited transformation.
09 10 – Transformation successful.
09 11 – BL21-DE3 transformed with Cd21L27+pCola did not grow.
09 12 – Too much bacteria grew up.
09 13 – Transformation unsuccessful. In some dishes still too many bacteria grown.
09 14 – Transformation unsuccessful.
09 15 – Transformation unsuccessful. Probably not enough pCDF-SP1 and pCDF-SP3 vector.
09 17 - Transformation successful.
About
| Date | Materials | Experiment |
|---|---|---|
| 08 26 |
P20 Carbicilin |
Transformation of P20 into JM109. |
| 08 27 |
P30 Chloramphenicol |
Transformation into JM109 (transformed with P20) |
| 08 28 |
JM109 transformants P30 Carbicilin Chloramphenicol |
Growing of JM109 transformants (low copy) and preparing for new transformation. Transformation of P30 into empty JM109. |
| 08 29 |
JM109 transformants P30 Carbicilin Chloramphenicol |
Transformation of P30 into JM109 (transformed with P20) |
| 08 30 |
JM109 transformants Carbicilin Chloramphenicol |
Transformation of P20 into JM109 (transformed with P30) |
| 08 31 |
JM109 transformants P30 P20 Carbicilin Chloramphenicol |
Transformation of P20 into JM109 (transformed with P30). Transformation of P30 into JM109 (transformed with P20). |
| 09 01 |
JM109 transformants JM109 (fresh) P20 P30 Carbicilin Chloramphenicol |
Transformation of P20 and P30 into JM109 (fresh). Transformation of P20 into JM109 (transformed with P30). Transformation of P30 into JM109 (transformed with P20). |
| 09 02 |
JM109 transformants (with P20 and P30) IPTG Carbicilin Chloramphenicol |
Growing transformants for 4 hours in a liquid LB. Transfer transformants into LB with different IPTG concentrations and grow overnight. |
| 09 12 | JM109 transformants (with P20 and P30) | Fluorescence measurement and analysis. |
| 09 06 |
JM109 transformants (with P20 and P30) IPTG Carbicilin Chloramphenicol |
Growing transformants for 4 hours in a liquid LB. Transfer transformants into LB with IPTG and grow overnight. |
| 09 07 | JM109 transformants (with P20 and P30) | Fluorescence measurement and analysis. |
| 09 08 |
JM109 transformants (with P20 and P30) IPTG Carbicilin Chloramphenicol |
Growing transformants for 4 hours in a liquid LB. Transfer transformants into LB with IPTG and grow overnight. |
| 09 10 | JM109 transformants (with P20 and P30) | Fluorescence measurement and analysis. |
| 09 11 | JM109 transformants (with P20 and P30) | Fluorescence measurement and analysis. |
| 09 13 |
JM109 transformants (with P20 and P30) pCDF vector Carbicilin Streptomycin |
Growing JM109 transformants as competent cells. Restriction and ligation of P30 fragment into pCDF vector. Transformation of new P30 into JM109 (transformed with P20). |
| 09 14 |
JM109 transformants (with P20 and P30) Carbicilin Streptomycin |
Colony PCR. |
| 09 15 |
JM109 transformants (with P20 and P30) Carbicilin Streptomycin |
Inoculating bacteria for 4 hours. Inoculating overnight with IPTG. |
| 09 16 | JM109 transformants (with P20 and P30) | Measuring fluorescence. |
| 09 17 | JM109 transformants (with P20 and P30) | Measuring fluorescence. |
Conclusions and Results:
08 26 – Transformation successful.
08 27 – Transformation unsuccessful.
08 29 – JM109 grown successfully. Transformation of P30 into empty JM109 was successful.
08 30 – Transformation unsuccessful.
08 31 - Transformation unsuccessful. The samples could have been pollen.
09 01 – Transformation unsuccessful. Maybe it was too much antibiotic on a petri dish.
09 02 – Transformation successful.
09 03 – Bad fluorimeter options.
09 07 – No repeated samples were made.
09 11 – Fluorescence measurement seems successful.
09 12 – Mistake found. P30 and P20 fragments are in an uncompetable plasmids. Repeating whole experiment.
09 14 - Transformation successful. Colony PCR successful.
09 17 - Fluorescence measurements are successful and conclusive.