Difference between revisions of "Team:Amsterdam//Project/Synthetic biology/Dependecies"
| Line 152: | Line 152: | ||
</div> | </div> | ||
| + | |||
| + | <div class="6u"> | ||
| + | <section class="special"> | ||
| + | |||
| + | <!-- the caption thingy, now you have to define the pictures in this way if you want the caption --> | ||
| + | <figure class = "image"> | ||
| + | <img src="https://upload.wikimedia.org/wikipedia/en/thumb/6/64/Urea_cycle_1.png/1024px-Urea_cycle_1.png" alt="<i>Synechocystis</i> interaction with <i>E. coli</i>"> | ||
| + | <figcaption> Figure 1. - Synechocystis Auxotroph being supplied nutrients from nutrient producing E. coli</figcaption> | ||
| + | </figure> | ||
| + | </section> | ||
| + | </div> | ||
</div> | </div> | ||
</div> | </div> | ||
Revision as of 22:02, 17 September 2015
Dependent Synechocystis
Some subtitle
Overview
Background
Background into the rational behind this module
Aim
Engineer an auxotrophic Synechocystis.
Methods
How to create an auxotrophic Synechocystis
Results
What was achieved
Parts
List of created parts.
Background
One can construct a synthetic consortium between two species with a variety of methods. Synechocystis could simply be engineered to export a carbon source (such as glucose) for E. coli in a glucose-absent medium. Although such a commensal relationship would definitely under certain conditions be considered stable, E. coli alone is the benefactor - there is no drive for Synechocystis to produce this carbon source. Increased stability is possible by creating a reciprocal relationship between the two species. Therefore, how can Cyanobacteria benefit or depend on E. coli’s presence? Cyanobacteria manufacturers it’s own food and has grown for centuries requiring only sunlight and water, in addition to inorganic elements. In fact, E. coli’s presence actually harms Cyanobacteria by limiting access to precious sunlight. Herein lies this module’s focus: Genetic engineering of an E. Coli dependent Synechocystis for the formation of stable synthetic consortia.
A basic requirement of consortia requires ‘communication’ between the species. Here we will attempt to construct a mutualistic relationship in which survival of one species is obligatory and beneficial for the survival of the other. The mode of communication will be the mutual production and exchange of essential metabolites. Since Synechocystis is an autotroph, one way to realize such a co-dependency is through the engineering of an auxotrophic bacterial strain. E. coli, could then in exchange produce the nutrient Synechocystis has been engineered to require. Ultimately, Synechocystis will produce a carbon source necessary for survival of E. coli, while E. coli produces a nutrient necessary for survival of Synechocystis - an engineered interdependence pathway.

Aim
This module will aim to engineer an auxotrophic Synechocystis. It also aims to test this auxotroph in the presence of an E. coli engineered to produce the nutrient that Synechocystis has been engineered to be deficient in. This is visualized in Figure 1. One might ponder what is the added benefit of this feedback system in our consortia. Indeed, as stated before, stability can be achieved with just Synechocystis feeding E. coli. However, with this feedback loop, we gain numerous benefits over the aforementioned system.
- We consider potential outbreak risks. An E.coli dependent Synechocystis would not be able to survive outside the lab, thus the risk of environmental contamination would diminish.
- An essential part of our project involves the creation of an emulsion based protocol to test potential consortia. If Synechocystis did not require E. coli, testing out the most effective and stable consortia in this manner would result in simply selecting against the presence of E. coli, as Synechocystis would do whatever it takes to increase it’s own growth rate. By having a feedback loop, we are therefore able to select consortia in which the members work well together.
- Assuming that E. coli could produce Arginine faster than Synechocystis can itself. By knocking out the gene to encode for the enzyme to produce arginine, Synechocystis does not need to waste energy in making these enzymes.
Auxotroph Criteria
There are several criteria that to be upheld when making the choice of what sort of auxotroph should be made. In our modeling attempts, an algorithm was developed to searched the metabolic map of Synechocystis and output candidate nutrients that fulfilled the following criteria.
- Simple. We want to choose a nutrient that can be relatively simple to knock out. Thus in our search, we focused on nutrients that only required the knock out of one gene to produce the desired phenotype.
- Loss of the gene encoding the reaction to produce the nutrient would result in non-growth. If the nutrient is not in the medium, Synechocystis should not grow. Reintroduction of the nutrient to the medium would result in growth of Synechocystis.
- The nutrient should be able to be produced and secreted by E. coli.
Auxotroph Targets
With the aforementioned criteria in mind, the auxotroph finder that was developed by one of our team members was able to find a variety of different suitable nutrients for which would be suitable candidates. Of these, two amino acids were chosen to be knocked out: Arginine and Proline.
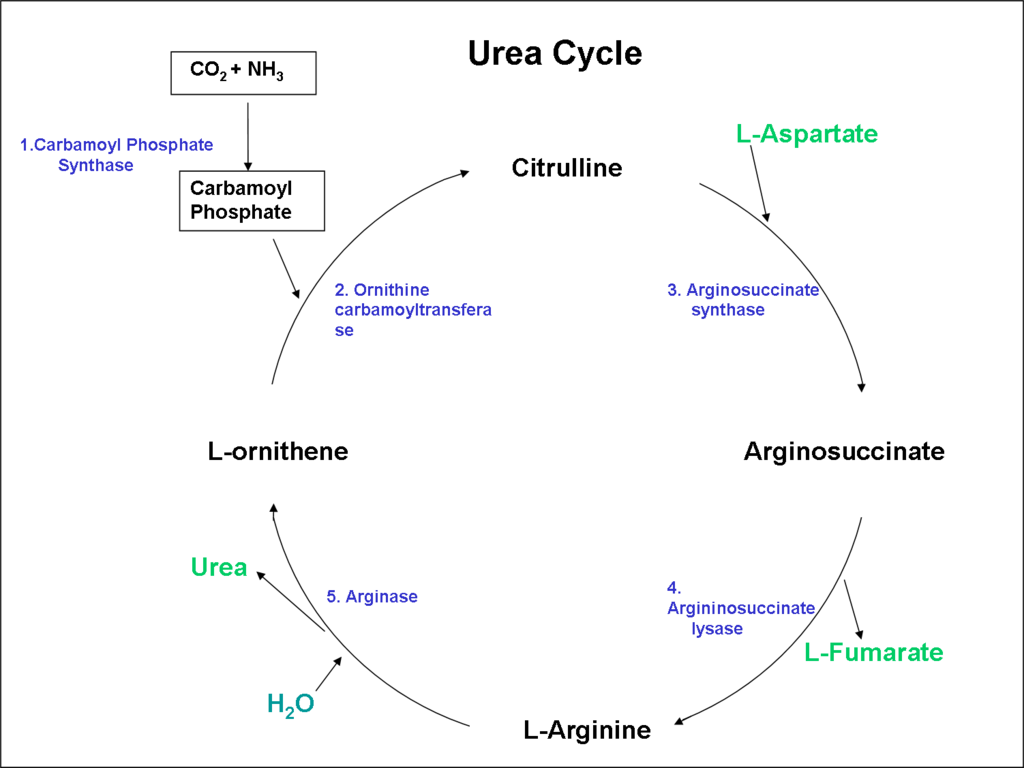
Arginine was a suitable target as we found a strain of E. coli that was able to synthesize it. In addition it has been shown that Synechocystis cells can grow faster in the presence of Arginine. The enzyme responsible for Arginine production in Synechocystis is L-arginosuccinate lyase (ASL) enzyme. This is encoded by the slr1133: ArgH gene. ASL drives the reaction from argininosuccinate into arginine and fumarate in the Urea Cycle. Figure 2 displays the mechanism behind Arginine production.
In the case of Proline, a strain of Salmonella and E. coli capable of producing Proline was procured. The enzyme responsible for Proline production in Synechocystis is pyrroline-5-carboxylate reductase. This is encoded by the slr0661: ProC gene. Figure three displays the mechanism behind Proline production in Synechocystis.