Difference between revisions of "Team:Heidelberg/Overview"
| Line 6: | Line 6: | ||
<div class="container"> | <div class="container"> | ||
<div class="content"> | <div class="content"> | ||
| − | <img src="https://static.igem.org/mediawiki/2015/ | + | <img src="https://static.igem.org/mediawiki/2015/2/26/Heidelberg_media_banner_overview.svg" style="width:100%; top:70px; margin-top: -10px; margin-left: -20px;"> |
</div> | </div> | ||
</div> | </div> | ||
Revision as of 22:39, 17 September 2015

Diving into the world of functional nucleic acids
In 1952 DNA was described as the carrier of genetic information for the first time.
Catalytic RNA – Ribozymes

The secondary structure of a Hammerhead Ribozyme (HHR). Arrow indicates position of cleavage between third and first stem. First and second stem are stem loops while stem three is open.
In 1982 the first catalytic RNA (Ribozyme): a self-spicing intron from Tetrahymena pre-rRNA was described.
Another ribozyme that is related to the HHR is the hepatitis δ virus ribozyme (HDV).

Naturally occurring Hairpin Ribozyme with cleavage site indicated by arrow.
Both of the mentioned types of ribozymes are found in satellite RNA of plant origin
Our idea of working with functional nucleic acids originated from this system. We were fascinated by the vast variety of processes that they can perform and started digging deeper into the potential of nucleic acids as tools. During this process we came across interesting systems amongst which we found the twin ribozyme (Fig. 3). Another famous hairpin-ribozyme-derived functional nucleic acid developed by Müller.
Next to the directed evolution of existing ribozymes in vitro selection methods

Secondary structure of the Twin Ribozyme. With cleavage and ligation sites indicated by arrows. The Hairpin Ribozyme is derived from the Hairpin Ribozyme.
Aptamers
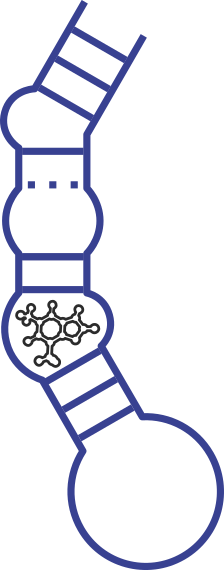
Aptamers are RNA or DNA sequences that can selectively bind to a ligand or target. Scientists have selected a variety of aptamers for small molecules ranging from theophylline (Fig. 4A) being one of the first described aptamers
Both catalytic nucleic acids and aptamers are as for today selected via systematic evolution of ligands by exponential enhancement
It has been shown that selected aptamers can be used to make the catalytic activity dependent on the presence of a ligand.

An aptamer connected via a communication module to an catalytic sequence; The communication module translates the interaction of the ligand with the aptamer to the catalytic part of the aptazyme.

Systematic Evolution of Ligands by EXponential Enrichment (SELEX): Current method to select functional nucleic acids from a random library. Figure adapted from: Source

The Spinach Aptamer turns on fluorescence of its ligand upon bind it.
Besides the above describes functional nucleic acids SELEX revealed aptamers which not only bind to a ligand but also are able to turn on fluorescence of different non-fluorescent dyes. An interesting example for an aptamer with such a special function is the Malachite Green Aptamer.
Though we so far mainly discussed functional RNA there are also both DNA aptamers and catalytic DNA which were selected using SELEX as well. As for today the concept of functional nucleic acids in nature has only been found for RNA. Nevertheless DNA aptamers and also catalytic DNA were selected in vitro.
Catalytic DNA – DNAzymes

Secondary structure of 7-18 and 10-23 RNA-cleaving DNAzyme with substrate RNA. Cleavage site is indiacted with an arrow.
SELEX revealed a variety of DNA aptamers and catalytic DNA known as DNAzymes (or Deoxribozymes). Breaker and Joyce selected the first DNAzyme, a RNA-cleaving DNA.
To verify the prediction of the software MAWS we fused the calculated aptamers to a DNAzyme with JAWS and thus created a ligand dependent activity. For this we used the HRP-mimicking DNAzyme (Fig. 9) which was described in 1998.