Team:TJU/Future Work

Although we made several innovative research in our project, there are still many other aspects to develop and optimize in the future. 1. We plan to find out biofilm parts of E.coli and modify several key point to reduce adhesion effect. 2. We need to further adjust ratio of our three-strain system to bring a quicker speed toward maximum. 3. Keeping power generation unchanged, we can reduce the volume of our device so that it will increase current density. 4. Making improvements in Shewanella for more electricity output. We intend to weaken any other unrelated functions except for power production in Shewanella and using co-culture system to produce necessary nutrition for its survival.
Proteolysis system
During the experiment, our MFC in anaerobic condition has been faced with unexpected lower pH harmful to microbe consortia. We aim to construct a more stable and robust MFC system, which will achieve the self-regulating and more sustainable output without excessive manipulation. So we have done some research on the acid-sensing and proteolysis system in bacteria. Based on the research, we attempt to build up “Sensing-Regulating” system. Our research and preliminary are as follows.
1 Sensing system—sensing the fluctuation of key metabolites
Every genus of cell has an optimum range of pH to live and Shewanella along with E. coliis no exception. Previous research has identified that Shewanella can maintain a maximal population density when pH keeps above 6.2 while E. colican survive in 5.6~8.1.[1] As an attempt to develop the superior sensing mechanism, we screen out promoter 170 (P170) and regulatory protein RcfB and introduce them into E. colifor adaption.
P170, a strongly acid-inducible promoter from Lactococcus lactis, is up-regulated at low pH during the transition to stationary phase.[2] The trans-acting protein RcfB, however, is involved in basal activity of P170 and is also essential for pH induction. The protein RcfB, upon activation by lactate, bind a DNA recognition motif within a promoter region and activates transcription.[3] When the cells are exposed to acid environments, the RcfB activation by an ‘acid’ signal allows its binding to the ACiD-box, resulting in transcription activation.[3]
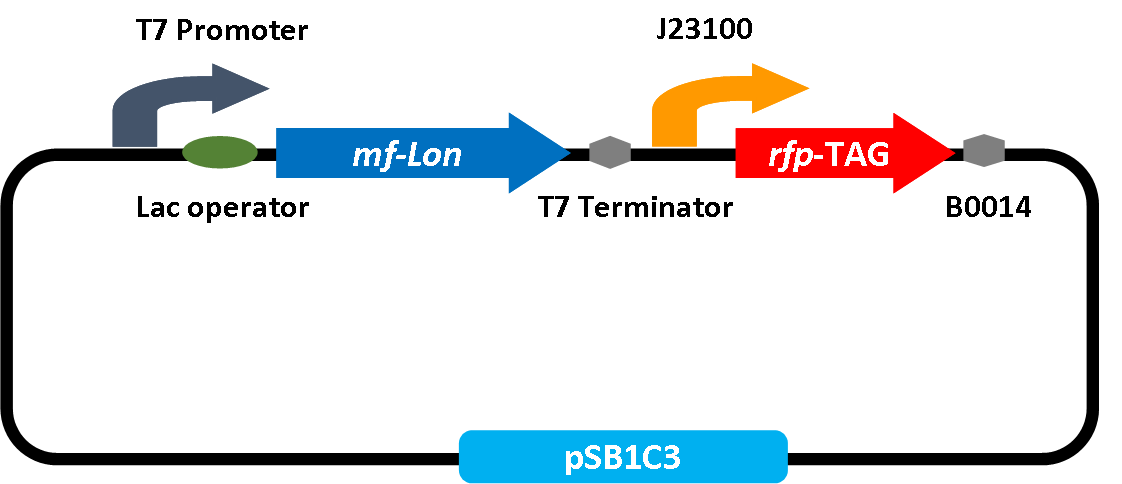
To test the idea that P170 is sensitive to acidity under the regulation of RcfB, we construct RFP gene into circuit containing P170 and RcfB. The RFP is expressed on the basis of P170 activation and RcfB which functions as a novel regulatory protein however, is controlled by a constitutive promoter J23100. High level of P170 activity required both RcfB and acidic conditions. Therefore, with the constitutive expression of RcfB, once the pH reaches the threshold that usually ranges from pH 6.0 to pH 6.5, P170 will be induced so that RFP can reflect red fluorescence under UV detection. Although RFP verification test is well developed, other functional genes also replace RFP gene as a strategy for application extension. In our later experiments, we substitute RFP for T7 polymerase for regulatory purpose.
2 Regulating
After activated sensing system which determines the pH threshold to suppress bacterial population density, we need to figure out how to retrieve to the proper acidity for living. Here comes our regulating system for acid recovery.
The ssrA tag sequence (mf-ssrA tag) and the Lon protease (mf-Lon) from Mesoplasma florum are made up of the crucial elements of a protein degradation system. Experiments has been identified that mf-ssrA tag is efficiently recognized by mf-Lon where the tag appends to C terminus of native or denatured proteins resulted in their rapid proteolysis by mf-Lon.[4] Besides, degradation system based on the Gram-positive M. florum tmRNA system does not rely on host degradation systems and can function in a wide range of bacteria, which makes it an adaptive method in variety. Here, we use this proteolysis system for lactate regulation and keep it in a oscillatory value of acidity.
Specifically, we utilize the effect of mf-Lon as well as ldhE bearing with tag under the control T7 promoter and J23100 promoter respectively, to degrade the overmuch lactate. We also aware that T7 promoter activation requires the combination of T7 RNA polymerase which is extremely promoter-specific and transcribes only DNA downstream of a T7 promoter.[5] According to the extensive study of our ‘sensing’ system, we substitute the RFP for T7 polymerase hoping that lower pH stage can activate T7 polymerase activation which then combines with T7 promoter for acting and mf-Lon, in this way, also expresses. It turns out that ldhE togethered with mf-ssrA tag can be degraded by cytoplasmic mf-Lon. So the pH can maintain in a proper level for cells’ living.
(实验表征结果和说明)Notably, the proteolysis system has potential to apply in many fields. As a strategy to make it a toolbox for further research, we use different tags to test the strength of its proteolysis. With the reflection of red fluorescence, we are able to detect mf-Lon activity on various tags based on the distinguish value of UV detection. This toolbox is intended to be a fast and convenient method for prospective proteolysis option and degradation research.
(实验表征结果和说明)References
[1] Min C, Chung H, Choi W, et al. Linear correlation between inactivation of E. coli and OH radical concentration in TiO2 photocatalytic disinfection.[J]. Water Research, 2004, 38(4):1069–1077. [2] Madsen S M, Arnau J, Vrang A, et al. Molecular characterization of the pH-inducible and growth phase-dependent promoter P170 of Lactococcus lactis.[J]. Molecular Microbiology, 1999, 32(1):75-87. [3] Madsen, Søren M, Hindré, Thomas, Le Pennec, Jean‐Paul, et al. Two acid‐inducible promoters from Lactococcus lactis require the cis‐acting ACiD‐box and the transcription regulator RcfB[J]. Molecular Microbiology, 2005, 56(3):735-746. [4] Eyal G, Sauer R T. Evolution of the ssrA degradation tag in Mycoplasma: specificity switch to a different protease.[J]. Proceedings of the National Academy of Sciences, 2008, 105(42):16113-16118. [5] Martin C T, Esposito E A, Theis K, et al. Structure and function in promoter escape by T7 RNA polymerase.[J]. Prog Nucleic Acid Res Mol Biol, 2005, 80(80):323–347.
E-mail: ggjyliuyue@gmail.com |Address: Building No.20, No.92 Weijin road, Tianjin University, China | Zip-cod: 300072
Copyright 2015@TJU iGEM Team